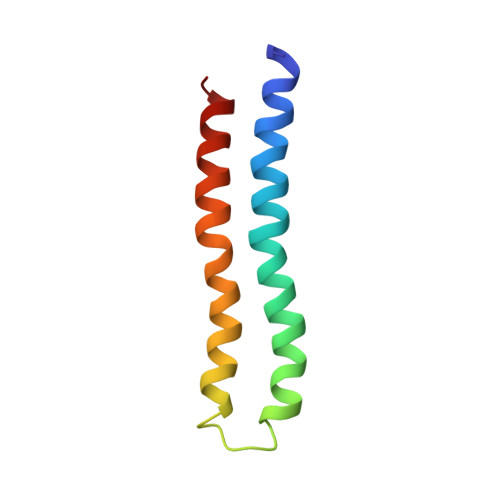
Solution structure of the transmembrane H+-transporting subunit c of the F1F0 ATP synthase.
Girvin, M.E., Rastogi, V.K., Abildgaard, F., Markley, J.L., Fillingame, R.H.(1998) Biochemistry 37: 8817-8824
- PubMed: 9636021 Search on PubMed
- DOI: https://doi.org/10.1021/bi980511m
- Primary Citation Related Structures:
1A91, 1C0V - PubMed Abstract:
Subunit c is the H+-translocating component of the F1F0 ATP synthase complex. H+ transport is coupled to conformational changes that ultimately lead to ATP synthesis by the enzyme. The properties of the monomeric subunit in a single-phase solution of chloroform-methanol-water (4:4:1) have been shown to mimic those of the protein in the native complex. Triple resonance NMR experiments were used to determine the complete structure of monomeric subunit c in this solvent mixture. The structure of the protein was defined by >2000 interproton distances, 64 (3)JN alpha, and 43 hydrogen-bonding NMR-derived restraints. The root mean squared deviation for the backbone atoms of the two transmembrane helices was 0.63 A. The protein folds as a hairpin of two antiparallel helical segments, connected by a short structured loop. The conserved Arg41-Gln42-Pro43 form the top of this loop. The essential H+-transporting Asp61 residue is located at a slight break in the middle of the C-terminal helix, just prior to Pro64. The C-terminal helix changes direction by 30 +/- 5 degrees at the conserved Pro64. In its protonated form, the Asp61 lies in a cavity created by the absence of side chains at Gly23 and Gly27 in the N-terminal helix. The shape and charge distribution of the molecular surface of the monomeric protein suggest a packing arrangement for the oligomeric protein in the F0 complex, with the front face of one monomer packing favorably against the back face of a second monomer. The packing suggests that the proton (cation) binding site lies between packed pairs of adjacent subunit c.
- Department of Biomolecular Chemistry, University of Wisconsin Medical School, Madison 53706, USA. girvin@aecom.yu.edu
Organizational Affiliation: