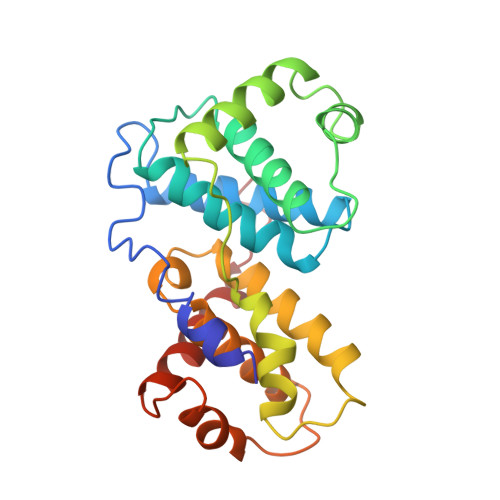
Crystal structure of a viral cyclin, a positive regulator of cyclin-dependent kinase 6.
Schulze-Gahmen, U., Jung, J.U., Kim, S.H.(1999) Structure 7: 245-254
- PubMed: 10368294 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(99)80035-5
- Primary Citation Related Structures:
1BU2 - PubMed Abstract:
Cyclin-dependent kinases (CDKs) have a central role in cell-cycle control and are activated by complex formation with positive regulatory proteins called cyclins and by phosphorylation. The overexpression and mutation of cyclins and CDKs has been associated with tumorigenesis and oncogenesis. A virus-encoded cyclin (v-cyclin) from herpesvirus saimiri has been shown to exhibit highest sequence homology to type D cyclins and specifically activates CDK6 of host cells to a very high degree. We have determined the first X-ray structure of a v-cyclin to 3.0 A resolution. The structure of the core domains is very similar to those of cyclin A and cyclin H from human cells. To understand the structural basis for the v-cyclin specificity for CDK6 and the insensitivity of the complex to inhibitors of the p21 and INK4 families, a v-cyclin-CDK2 model was built on the basis of the known structures of human cyclin A in complex with CDK2 and the CDK inhibitor p27(Kip1). Although many critical interactions between cyclin A and CDK2 would be conserved in a v-cyclin-CDK2 complex, some appear sterically or electrostatically unfavorable due to shifts in the backbone conformation or sidechain differences and may contribute to v-cyclin selectivity for CDK6. The insensitivity of v-cyclin-CDK6 complexes to inhibitors of the p21 family is probably due to structural changes in v-cyclin that lead to a flatter surface area offering fewer potential contacts with the protein inhibitor. In addition, sequence changes in v-cyclin eliminate hydrogen-bonding partners for atoms of the p27(Kip1) inhibitor. This structure provides the first model for interactions between v-cyclins and host cell-cycle proteins; these interactions may be important for virus survival as well as oncogenic transformation of host cells.
- Department of Chemistry, Earnest Orlando Berkeley National Laboratory, University of California, Berkeley, California, 94720 USA. uschulze-gahmen@lbl.gov
Organizational Affiliation: