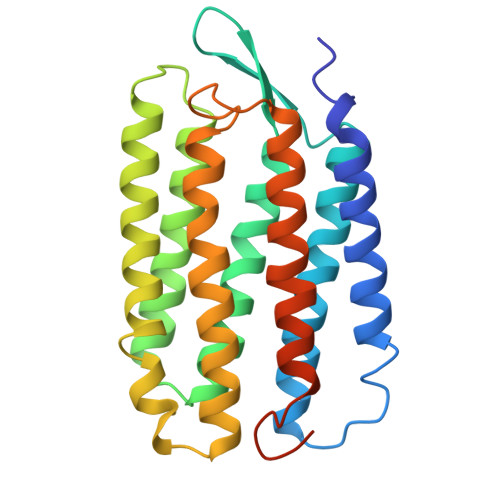
Lipid patches in membrane protein oligomers: crystal structure of the bacteriorhodopsin-lipid complex
Essen, L., Siegert, R., Lehmann, W.D., Oesterhelt, D.(1998) Proc Natl Acad Sci U S A 95: 11673-11678
- PubMed: 9751724
- DOI: https://doi.org/10.1073/pnas.95.20.11673
- Primary Citation of Related Structures:
1BRR - PubMed Abstract:
Heterogenous nucleation on small molecule crystals causes a monoclinic crystal form of bacteriorhodopsin (BR) in which trimers of this membrane protein pack differently than in native purple membranes. Analysis of single crystals by nano-electrospray ionization-mass spectrometry demonstrated a preservation of the purple membrane lipid composition in these BR crystals. The 2.9-A x-ray structure shows a lipid-mediated stabilization of BR trimers where the glycolipid S-TGA-1 binds into the central compartment of BR trimers. The BR trimer/lipid complex provides an example of local membrane thinning as the lipid head-group boundary of the central lipid patch is shifted by 5 A toward the membrane center. Nonbiased electron density maps reveal structural differences to previously reported BR structures, especially for the cytosolic EF loop and the proton exit pathway. The terminal proton release complex now comprises an E194-E204 dyad as a diffuse proton buffer.
- Max-Planck-Institut fur Biochemie, Am Klopferspitz 18a, D-82152 Martinsried, Germany.
Organizational Affiliation: