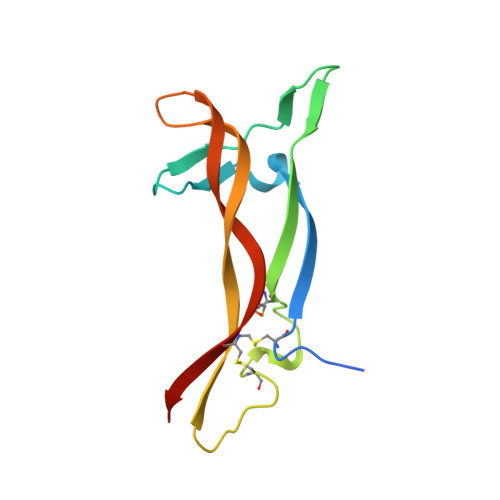
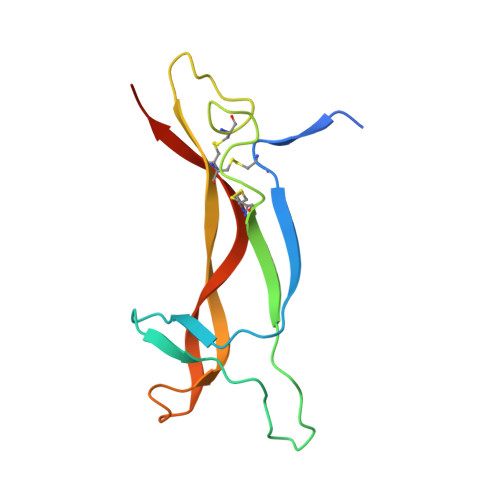
Structure of the brain-derived neurotrophic factor/neurotrophin 3 heterodimer.
Robinson, R.C., Radziejewski, C., Stuart, D.I., Jones, E.Y.(1995) Biochemistry 34: 4139-4146
- PubMed: 7703225 Search on PubMed
- DOI: https://doi.org/10.1021/bi00013a001
- Primary Citation Related Structures:
1BND - PubMed Abstract:
The development and sustenance of specific neuronal populations in the peripheral and central nervous systems are controlled through the binding of neurotrophic factors to high-affinity cell surface receptors. The neurotrophins (nerve growth factor, NGF; brain-derived neurotrophic factor, BDNF; neurotrophin 3, NT3; and neurotrophin 4, NT4) are dimeric molecules which share approximately 50% sequence identity. The crystal structure of the murine NGF homodimer [McDonald et al. (1991) Nature 354, 411-414] indicated that the dimer interface corresponds to regions of high sequence conservation throughout the neurotrophin family. This potential compatibility was duly exploited for the production in vitro of noncovalent heterodimers between the different neurotrophins [Radziejewski, C., & Robinson, R.C. (1993) Biochemistry 32, 13350-13356; Jungbluth et al. (1994) Eur. J. Biochem. 221, 677-685]. Here, we report the X-ray structure at 2.3 A resolution of one such heterodimer, between human BDNF, and human NT3. The NGF, BDNF, and NT3 protomers share the same topology and are structurally equivalent in regions which contribute to the dimer interface in line with the propensity of the neurotrophins to form heterodimers. Analysis of the structure of regions of the BDNF/NT3 heterodimer involved in receptor specificity led us to conclude that heterodimer binding to p75 involves distant binding sites separately located on each protomer of the heterodimer. In contrast, heterodimer interactions with the trk receptors probably utilize hybrid binding sites comprised of residues contributed by both protomers in the heterodimer. The existence of such hybrid binding sites for the trk receptor provides an explanation for the lower activity of the BDNF/NT3 heterodimer in comparison to the homodimers.(ABSTRACT TRUNCATED AT 250 WORDS)
- Laboratory of Molecular Biophysics, Oxford University, United Kingdom.
Organizational Affiliation: