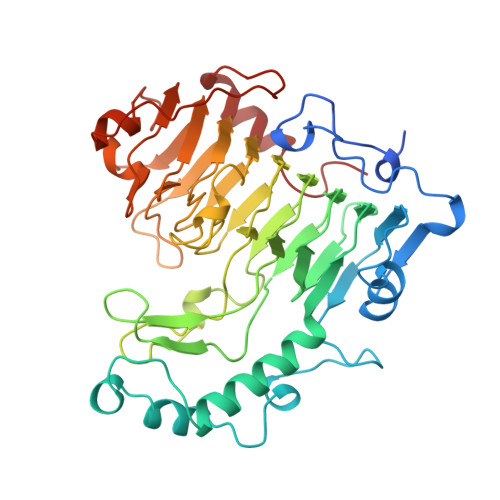
The structure of Bacillus subtilis pectate lyase in complex with calcium.
Pickersgill, R., Jenkins, J., Harris, G., Nasser, W., Robert-Baudouy, J.(1994) Nat Struct Biol 1: 717-723
- PubMed: 7634076
- DOI: https://doi.org/10.1038/nsb1094-717
- Primary Citation of Related Structures:
1BN8 - PubMed Abstract:
We have solved the structure of the Bacillus subtilis pectate lyase (BsPel) in complex with calcium. The structure consists of a parallel beta-helix domain and a loop region. The alpha L-bounded beta-strand seen in BsPel is a new element of protein structure and its frequent occurrence suggests it is an important characteristic of the parallel beta-helix. A pronounced cleft is formed between the loops and the parallel beta-helix domain and we propose that this is the active site cleft. Calcium, essential for the activity of the enzyme, binds at the bottom of this cleft and an arginine residue close to the calcium, which is conserved across all pectin and pectate lyases, may be involved in catalysis.
- Department of Protein Engineering, Institute of Food Research, Reading, UK.
Organizational Affiliation: