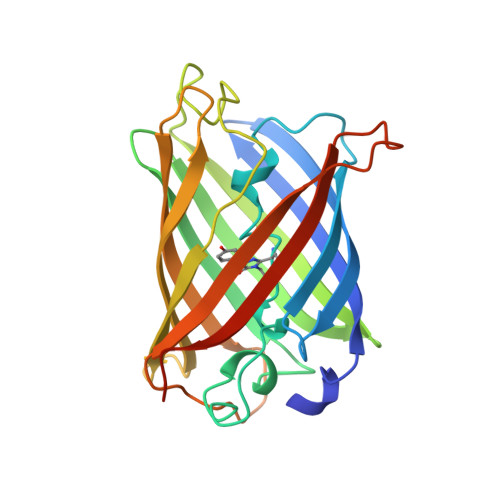
Crystal structure and refolding properties of the mutant F99S/M153T/V163A of the green fluorescent protein.
Battistutta, R., Negro, A., Zanotti, G.(2000) Proteins 41: 429-437
- PubMed: 11056031 Search on PubMed
- DOI: https://doi.org/10.1002/1097-0134(20001201)41:4<429::aid-prot10>3.0.co;2-d
- Primary Citation Related Structures:
1B9C - PubMed Abstract:
The mutant F99S/M153T/V163A of the Green Fluorescent Protein (c3-GFP) has spectral characteristics similar to the wild-type GFP, but it is 42-fold more fluorescent in vivo. Here, we report the crystal structure and the refolding properties of c3-GFP and compare them with those of the less fluorescent wt-GFP and S65T mutant. The topology and the overall structure of c3-GFP is similar to the wild-type GFP. The three mutated residues, Ser99, Thr153, and Ala163, lie on the surface of the protein in three different beta-strands. The side chains of Ser99 and Thr153 are exposed to the solvent, whereas that of Ala163 points toward the interior of the protein. No significant deviation from the structure of the wild-type molecule is found around these positions, and there is not clear evidence of any distortion in the position of the chromophore or of the surrounding residues induced by the mutated amino acids. In vitro refolding experiments on urea-denatured c3-GFP reveal a renaturation behavior similar to that of the S65T molecule, with kinetic constants of the same order of magnitude. We conclude that the higher fluorescence activity of c3-GFP can be attributed neither to particular structural features nor to a faster folding process, as previously proposed.
- Dipartimento di Chimica Organica and Centro Studi sui Biopolimeri, Università di Padova, Padova, Italy. roberto@chor.unipd.it
Organizational Affiliation: