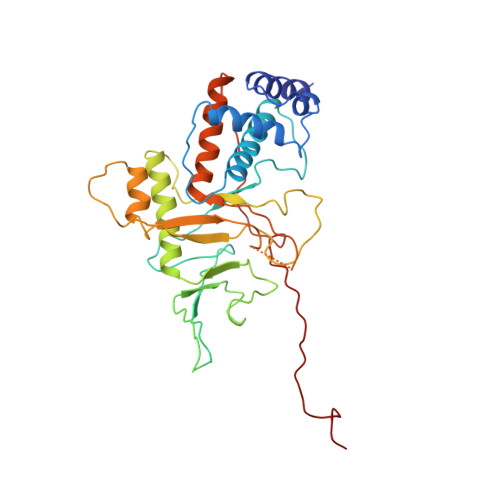
Conformational change of the adenovirus DNA-binding protein induced by soaking crystals with K3UO2F5 solutions.
Kanellopoulos, P.N., Tsernoglou, D., van der Vliet, P.C., Tucker, P.A.(1996) Acta Crystallogr D Biol Crystallogr 52: 942-945
- PubMed: 15299602 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444996005525
- Primary Citation Related Structures:
1ANV - PubMed Abstract:
Soaking crystals of the C-terminal DNA-binding domain of the adenovirus single-stranded DNA-binding protein with a buffer containing K(3)UO(2)F(5) results in a 9% change of the crystallographic c axis without destruction of the crystals or appreciable loss of resolution. The crystals belong to space group P2(1)2(1)2(1) with a = 79.7, b = 75.6 and c = 60.6 A. The three-dimensional structure has been refined to 2.7 A with a crystallographic R factor of 0.206. Antiparallel chains of protein molecules running through the entire crystal are linked by uranyl ions. The relative orientation of protein monomers is flexible, even in the crystalline state, and allows changes in the packing of the protein chains.
- Structural Biology Programme, European Molecular Biology Laboratory, Heidelberg, Germany.
Organizational Affiliation: