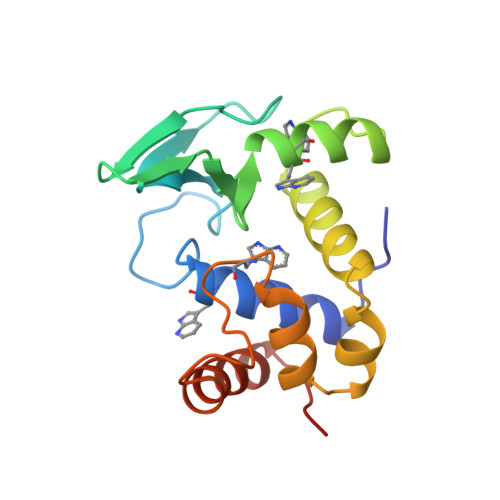
Crystal structure of the lysozyme from bacteriophage lambda and its relationship with V and C-type lysozymes.
Evrard, C., Fastrez, J., Declercq, J.P.(1998) J Mol Biology 276: 151-164
- PubMed: 9514719 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1997.1499
- Primary Citation Related Structures:
1AM7 - PubMed Abstract:
Like other lysozymes, the bacteriophage lambda lysozyme is involved in the digestion of bacterial walls. This enzyme is remarkable in that its mechanism of action is different from the classical lysozyme's mechanism. From the point of view of protein evolution, it shows features of lysozymes from different classes. The crystal structure of the enzyme in which all tryptophan residues have been replaced by aza-tryptophan has been solved by X-ray crystallography at 2.3 A using a combination of multiple isomorphous replacement, non-crystallographic symmetry averaging and density modification techniques. There are three molecules in the asymmetric unit. The characteristic structural elements of lysozymes are conserved: each molecule is organized in two domains connected by a helix and the essential catalytic residue (Glu19) is located in the depth of a cleft between the two domains. This cleft shows an open conformation in two of the independent molecules, while access to the cavity is much more restricted in the last one. A structural alignment with T4 lysozyme and hen egg white lysozyme allows us to superpose about 60 C alpha atoms with a rms distance close to 2 A. The best alignments concern the helix preceding the catalytic residue, some parts of the beta sheets and the helix joining the two domains. The results of sequence alignments with the V and C lysozymes, in which weak local similarities had been detected, are compared with the structural results.
- Laboratoire de Chimie Physique et de Cristallographie, Université Catholique de Louvain, Louvain-la-Neuve Belgium.
Organizational Affiliation: