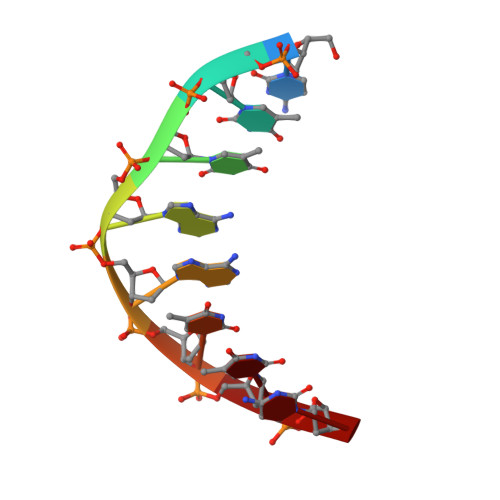
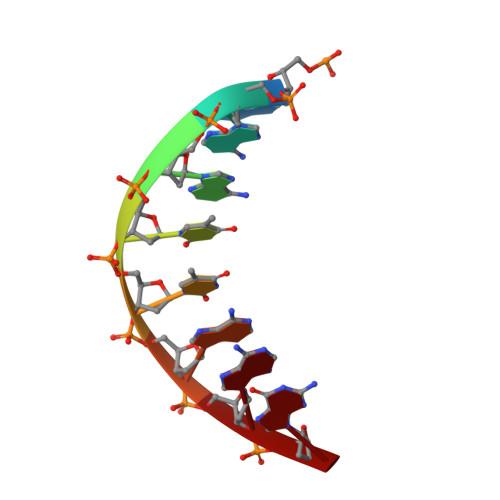
A host-guest approach for determining drug-DNA interactions: an example using netropsin.
Goodwin, K.D., Long, E.C., Georgiadis, M.M.(2005) Nucleic Acids Res 33: 4106-4116
- PubMed: 16049022 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki717
- Primary Citation Related Structures:
1ZTT, 1ZTW - PubMed Abstract:
Netropsin is a well-characterized DNA minor groove binding compound that serves as a model for the study of drug-DNA interactions. Our laboratory has developed a novel host-guest approach to study drug-DNA interactions in which the host, the N-terminal fragment of Moloney murine leukemia virus reverse transcriptase (MMLV RT) is co-crystallized with a DNA oligonucleotide guest in the presence and absence of drug. We have co-crystallized netropsin with the RT fragment bound to the symmetric 16mer d(CTTAATTCGAATTAAG)2 and determined the structure of the complex at 1.85 A. In contrast to previously reported netropsin-DNA structures, our oligonucleotide contains two AATT sites that bind netropsin with flanking 5' and 3' sequences that are not symmetric. The asymmetric unit of the RT fragment-DNA-netropsin crystals contains one protein molecule and one-half of the 16mer with one netropsin molecule bound. The guanidinium moiety of netropsin binds in a narrow part of the minor groove, while the amidinium is bound in the widest region within the site. We compare this structure to other Class I netropsin-DNA structures and find that the asymmetry of minor groove widths in the AATT site contributes to the orientation of netropsin within the groove while hydrogen bonding patterns vary in the different structures.
- Department of Biochemistry and Molecular Biology, Indiana University School of Medicine, IN 46202, USA.
Organizational Affiliation: