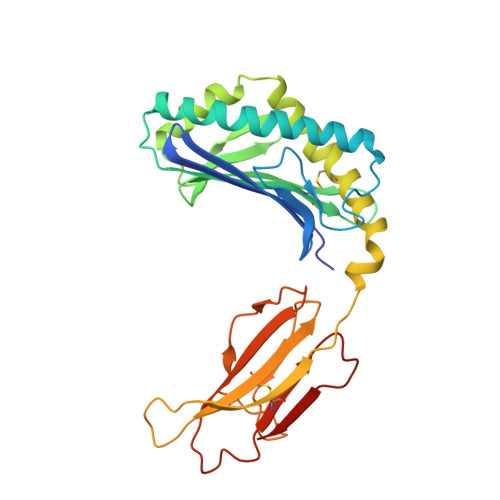
The crystal structure of human CD1d with and without alpha-galactosylceramide
Koch, M., Stronge, V.S., Shepherd, D., Gadola, S.D., Mathew, B., Ritter, G., Fersht, A.R., Besra, G.S., Schmidt, R.R., Jones, E.Y., Cerundolo, V.(2005) Nat Immunol 6: 819-826
- PubMed: 16007090 Search on PubMed
- DOI: https://doi.org/10.1038/ni1225
- Primary Citation Related Structures:
1ZT4 - PubMed Abstract:
The glycolipid alpha-galactosylceramide binds with high affinity to CD1d and stimulates natural killer T cells. Here we report the crystal structure of human CD1d in complex with synthetic alpha-galactosylceramide at a resolution of 3.0 A. The structure shows a tightly fit lipid in the CD1d binding groove, with the sphingosine chain bound in the C' pocket and the longer acyl chain anchored in the A' pocket. We also present the CD1d structure without lipid, which has a more open conformation of the binding groove, suggesting a dual conformation of CD1d in which the 'open' conformation is more able to load lipids. These structures provide clues as to how CD1 molecules load glycolipids as well as data to guide the design of new therapeutic agents.
- Cancer Research UK Receptor Structure Research Group, The Henry Wellcome Building for Genomic Medicine, Roosevelt Drive, Headington, Oxford OX3 7BN, UK.
Organizational Affiliation: