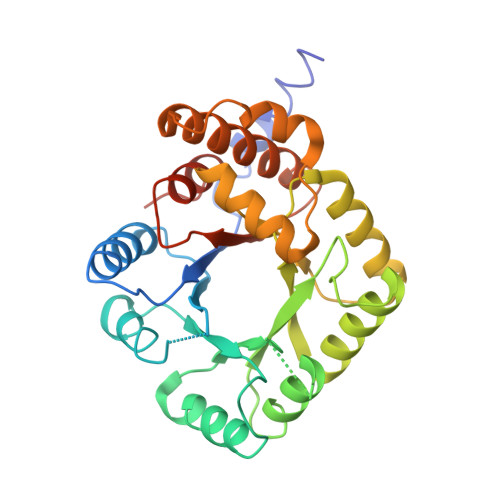
Structures of NADH and CH(3)-H(4)Folate Complexes of Escherichia coli Methylenetetrahydrofolate Reductase Reveal a Spartan Strategy for a Ping-Pong Reaction
Pejchal, R., Sargeant, R., Ludwig, M.L.(2005) Biochemistry 44: 11447-11457
- PubMed: 16114881 Search on PubMed
- DOI: https://doi.org/10.1021/bi050533q
- Primary Citation Related Structures:
1ZP3, 1ZP4, 1ZPT, 1ZRQ - PubMed Abstract:
Methylenetetrahydrofolate reductases (MTHFRs; EC 1.7.99.5) catalyze the NAD(P)H-dependent reduction of 5,10-methylenetetrahydrofolate (CH(2)-H(4)folate) to 5-methyltetrahydrofolate (CH(3)-H(4)folate) using flavin adenine dinucleotide (FAD) as a cofactor. The initial X-ray structure of Escherichia coli MTHFR revealed that this 33-kDa polypeptide is a (betaalpha)(8) barrel that aggregates to form an unusual tetramer with only 2-fold symmetry. Structures of reduced enzyme complexed with NADH and of oxidized Glu28Gln enzyme complexed with CH(3)-H(4)folate have now been determined at resolutions of 1.95 and 1.85 A, respectively. The NADH complex reveals a rare mode of dinucleotide binding; NADH adopts a hairpin conformation and is sandwiched between a conserved phenylalanine, Phe223, and the isoalloxazine ring of FAD. The nicotinamide of the bound pyridine nucleotide is stacked against the si face of the flavin ring with C4 adjoining the N5 of FAD, implying that this structure models a complex that is competent for hydride transfer. In the complex with CH(3)-H(4)folate, the pterin ring is also stacked against FAD in an orientation that is favorable for hydride transfer. Thus, the binding sites for the two substrates overlap, as expected for many enzymes that catalyze ping-pong reactions, and several invariant residues interact with both folate and pyridine nucleotide substrates. Comparisons of liganded and substrate-free structures reveal multiple conformations for the loops beta2-alpha2 (L2), beta3-alpha3 (L3), and beta4-alpha4 (L4) and suggest that motions of these loops facilitate the ping-pong reaction. In particular, the L4 loop adopts a "closed" conformation that allows Asp120 to hydrogen bond to the pterin ring in the folate complex but must move to an "open" conformation to allow NADH to bind.
- Department of Biological Chemistry and Biophysics Research Division, The University of Michigan, Ann Arbor, Michigan 48109-1055, USA.
Organizational Affiliation: