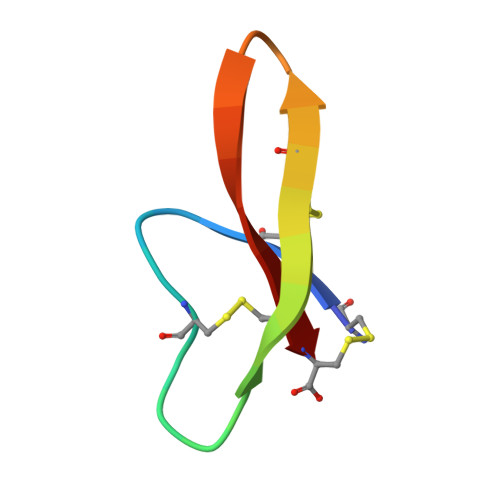
Reconstruction of the conserved beta-bulge in mammalian defensins using D-amino acids.
Xie, C., Prahl, A., Ericksen, B., Wu, Z., Zeng, P., Li, X., Lu, W.Y., Lubkowski, J., Lu, W.(2005) J Biological Chem 280: 32921-32929
- PubMed: 15894545 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M503084200
- Primary Citation Related Structures:
1ZMH, 1ZMI, 1ZMK - PubMed Abstract:
Defensins are cationic antimicrobial mini-proteins that play important roles in the innate immune defense against microbial infection. Six invariant Cys residues in each defensin form three structurally indispensable intramolecular disulfide bridges. The only other residue invariant in all known mammalian defensins is a Gly. Structural studies indicate that the invariant Gly residue is located in an atypical, classic-type beta-bulge with the backbone torsion angles (Phi, Psi) disallowed for L-amino acids but permissible for D-enantiomers. We replaced the invariant Gly17 residue in human neutrophil alpha-defensin 2 (HNP2) by L-Ala or one of the D-amino acids Ala, Glu, Phe, Arg, Thr, Val, or Tyr. Although L-Ala17-HNP2 could not be folded, resulting in massive aggregation, all of the D-amino acid-substituted analogs folded with high efficiency. The high resolution x-ray crystal structures of dimeric D-Ala17-HNP2 were determined in three different crystal forms, showing a well preserved beta-bulge identical to those found in other defensins. The seven D-analogs of HNP2 exhibited highly variable bactericidal activity against Gram-positive and Gram-negative test strains, consistent with the premise that interplay between charge and hydrophobicity dictates how amphiphilic defensins kill. Further, the bactericidal activity of these d-amino acid analogs of HNP2 correlated well with their ability to induce leakage from large unilamellar vesicles, supporting membrane permeabilization as the lethal event in microbial killing by HNP2. Our findings identify a conformational prerequisite in the beta-bulge of defensins essential for correct folding and native structure, thereby explaining the molecular basis of the Gly-Xaa-Cys motif conserved in all mammalian defensins.
- Institute of Human Virology, University of Maryland Biotechnology Institute, Baltimore, Maryland 21201, USA.
Organizational Affiliation: