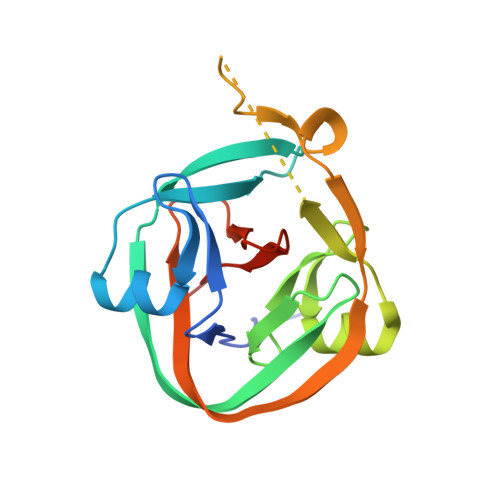
Crystal structures of an intein from the split dnaE gene of Synechocystis sp. PCC6803 reveal the catalytic model without the penultimate histidine and the mechanism of zinc ion inhibition of protein splicing
Sun, P., Ye, S., Ferrandon, S., Evans, T.C., Xu, M.Q., Rao, Z.(2005) J Mol Biology 353: 1093-1105
- PubMed: 16219320 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.09.039
- Primary Citation Related Structures:
1ZD7, 1ZDE - PubMed Abstract:
The first naturally occurring split intein was found in the dnaE gene of Synechocystis sp. PCC6803 and belongs to a subclass of inteins without a penultimate histidine residue. We describe two high-resolution crystal structures, one derived from an excised Ssp DnaE intein and the second from a splicing-deficient precursor protein. The X-ray structures indicate that His147 in the conserved block F activates the side-chain N(delta) atom of the intein C-terminal Asn159, leading to a nucleophilic attack on the peptide bond carbonyl carbon atom at the C-terminal splice site. In this process, Arg73 appears to stabilize the transition state by interacting with the carbonyl oxygen atom of the scissile bond. Arg73 also seems to substitute for the conserved penultimate histidine residue in the formation of an oxyanion hole, as previously identified in other inteins. The finding that the precursor structure contains a zinc ion chelating the highly conserved Cys160 and Asp140 reveals the structural basis of Zn2+-mediated inhibition of protein splicing. Furthermore, it is of interest to observe that the carbonyl carbon atom of Asn159 and N(eta) of Arg73 are 2.6 angstroms apart in the free intein structure and 10.6 angstroms apart in the precursor structure. The orientation change of the aromatic ring of Tyr-1 following the initial acyl shift may be a key switching event contributing to the alignment of Arg73 and the C-terminal scissile bond, and may explain the sequential reaction property of the Ssp DnaE intein.
- Laboratory of Structural Biology, Tsinghua University, Beijing 100084, People's Republic of China.
Organizational Affiliation: