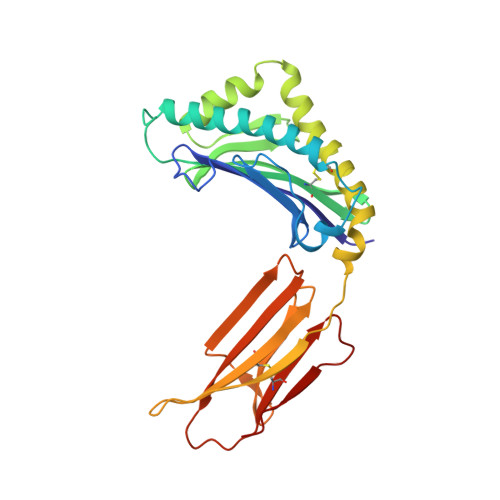
Crystal structure of human ZAG, a fat-depleting factor related to MHC molecules.
Sanchez, L.M., Chirino, A.J., Bjorkman, P.(1999) Science 283: 1914-1919
- PubMed: 10206894 Search on PubMed
- DOI: https://doi.org/10.1126/science.283.5409.1914
- Primary Citation Related Structures:
1ZAG - PubMed Abstract:
Zn-alpha2-glycoprotein (ZAG) is a soluble protein that is present in serum and other body fluids. ZAG stimulates lipid degradation in adipocytes and causes the extensive fat losses associated with some advanced cancers. The 2.8 angstrom crystal structure of ZAG resembles a class I major histocompatibility complex (MHC) heavy chain, but ZAG does not bind the class I light chain beta2-microglobulin. The ZAG structure includes a large groove analogous to class I MHC peptide binding grooves. Instead of a peptide, the ZAG groove contains a nonpeptidic compound that may be implicated in lipid catabolism under normal or pathological conditions.
- Division of Biology, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: