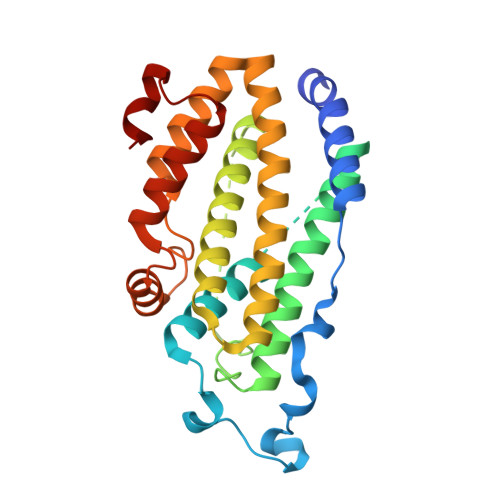
X-ray structure of putative acyl-ACP desaturase DesA2 from Mycobacterium tuberculosis H37Rv.
Dyer, D.H., Lyle, K.S., Rayment, I., Fox, B.G.(2005) Protein Sci 14: 1508-1517
- PubMed: 15929999 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.041288005
- Primary Citation Related Structures:
1ZA0 - PubMed Abstract:
Genome sequencing showed that two proteins in Mycobacterium tuberculosis H37Rv contain the metal binding motif (D/E)X(2)HX(approximately 100)(D/E)X(2)H characteristic of the soluble diiron enzyme superfamily. These putative acyl-ACP desaturase genes desA1 and desA2 were cloned from genomic DNA and expressed in Escherichia coli BL21(DE3). DesA1 was found to be insoluble, but in contrast, DesA2 was a soluble protein amenable to biophysical characterization. Here, we report the 2.0 A resolution X-ray structure of DesA2 determined by multiple anomalous dispersion (MAD) phasing from a Se-met derivative and refinement against diffraction data obtained on the native protein. The X-ray structure shows that DesA2 is a homodimeric protein with a four-helix bundle core flanked by five additional helices that overlay with 192 structurally equivalent amino acids in the structure of stearoyl-ACP Delta9 desaturase from castor plant with an rms difference 1.42 A. In the DesA2 crystals, one metal (likely Mn from the crystallization buffer) was bound in high occupancy at the B-site of the conserved metal binding motif, while the A-site was not occupied by a metal ion. Instead, the amino group of Lys-76 occupied this position. The relationships between DesA2 and known diiron enzymes are discussed.
- Department of Biochemistry, 433 Babcock Dr., University of Wisconsin, Madison, Madison, WI 53706, USA.
Organizational Affiliation: