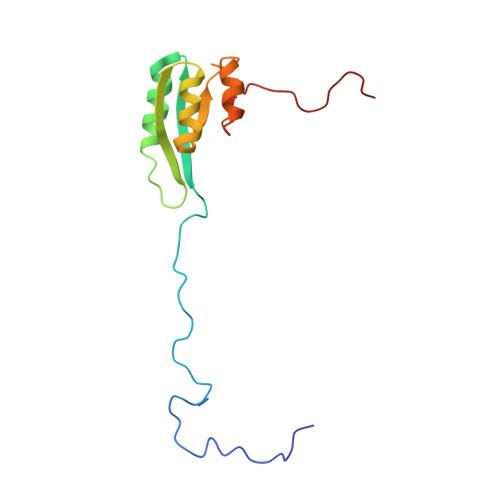
Solution structure of the C1-subdomain of Bacillus stearothermophilus translation initiation factor IF2
Wienk, H., Tomaselli, S., Bernard, C., Spurio, R., Picone, D., Gualerzi, C.O., Boelens, R.(2005) Protein Sci 14: 2461-2468
- PubMed: 16081655 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051531305
- Primary Citation Related Structures:
1Z9B - PubMed Abstract:
IF2 is one of three bacterial translation initiation factors that are conserved through all kingdoms of life. It binds the 30S and 50S ribosomal subunits, as well as fMet-tRNAf(Met). After these interactions, fMet-tRNAf(Met) is oriented to the ribosomal P-site where the first amino acid of the nascent polypeptide, formylmethionine, is presented. The C-terminal domain of Bacillus stearothermophilus IF2, which is responsible for recognition and binding of fMet-tRNAf(Met), contains two structured modules. Previously, the solution structure of the most C-terminal module, IF2-C2, has been elucidated by NMR spectroscopy and direct interactions between this subdomain and fMet-tRNAf(Met) were reported. In the present NMR study we have obtained the spectral assignment of the other module of the C-terminal domain (IF2-C1) and determined its solution structure and backbone dynamics. The IF2-C1 core forms a flattened fold consisting of a central four-stranded parallel beta-sheet flanked by three alpha-helices. Although its overall organization resembles that of subdomain III of the archaeal IF2-homolog eIF5B whose crystal structure had previously been reported, some differences of potential functional significance are evident.
- Bijvoet Center for Biomolecular Research, Department of NMR Spectroscopy, Utrecht University, Padualaan 8, 3584 CH Utrecht, The Netherlands.
Organizational Affiliation: