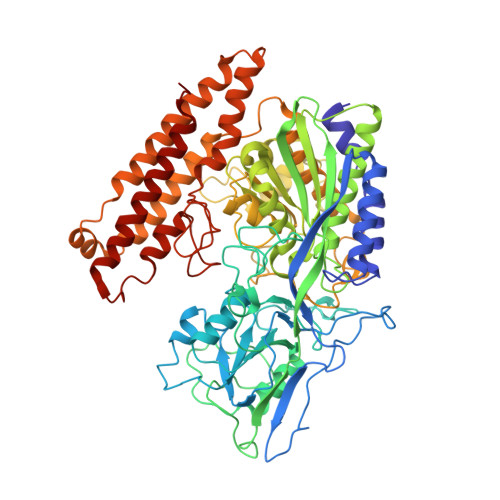
Crystal structure of prostate-specific membrane antigen, a tumor marker and peptidase
Davis, M.I., Bennett, M.J., Thomas, L.M., Bjorkman, P.J.(2005) Proc Natl Acad Sci U S A 102: 5981-5986
- PubMed: 15837926 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0502101102
- Primary Citation Related Structures:
1Z8L - PubMed Abstract:
Prostate-specific membrane antigen (PSMA) is highly expressed in prostate cancer cells and nonprostatic solid tumor neovasculature and is a target for anticancer imaging and therapeutic agents. PSMA acts as a glutamate carboxypeptidase (GCPII) on small molecule substrates, including folate, the anticancer drug methotrexate, and the neuropeptide N-acetyl-l-aspartyl-l-glutamate. Here we present the 3.5-A crystal structure of the PSMA ectodomain, which reveals a homodimer with structural similarity to transferrin receptor, a receptor for iron-loaded transferrin that lacks protease activity. Unlike transferrin receptor, the protease domain of PSMA contains a binuclear zinc site, catalytic residues, and a proposed substrate-binding arginine patch. Elucidation of the PSMA structure combined with docking studies and a proposed catalytic mechanism provides insight into the recognition of inhibitors and the natural substrate N-acetyl-l-aspartyl-l-glutamate. The PSMA structure will facilitate development of chemotherapeutics, cancer-imaging agents, and agents for treatment of neurological disorders.
- Division of Biology 114-96 and Howard Hughes Medical Institute, California Institute of Technology, Pasadena, CA 91125, USA.
Organizational Affiliation: