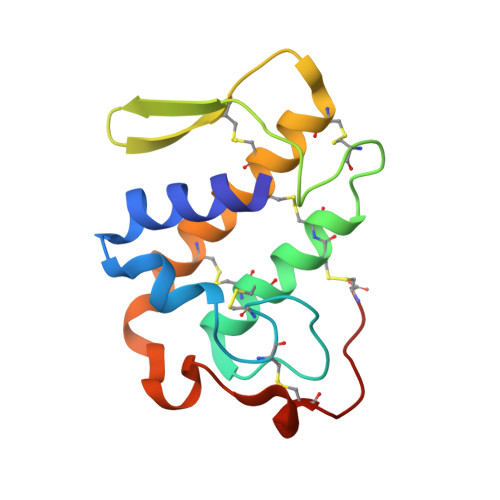
Structure of BthA-I complexed with p-bromophenacyl bromide: possible correlations with lack of pharmacological activity.
Magro, A.J., Takeda, A.A., Soares, A.M., Fontes, M.R.(2005) Acta Crystallogr D Biol Crystallogr 61: 1670-1677
- PubMed: 16301802 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905029598
- Primary Citation Related Structures:
1Z76 - PubMed Abstract:
The crystal structure of an acidic phospholipase A(2) isolated from Bothrops jararacussu venom (BthA-I) chemically modified with p-bromophenacyl bromide (BPB) has been determined at 1.85 Angstroms resolution. The catalytic, platelet-aggregation inhibition, anticoagulant and hypotensive activities of BthA-I are abolished by ligand binding. Electron-density maps permitted unambiguous identification of inhibitor covalently bound to His48 in the substrate-binding cleft. The BthA-I-BPB complex contains three structural regions that are modified after inhibitor binding: the Ca(2+)-binding loop, beta-wing and C-terminal regions. Comparison of BthA-I-BPB with two other BPB-inhibited PLA(2) structures suggests that in the absence of Na(+) ions at the Ca(2+)-binding loop, this loop and other regions of the PLA(2)s undergo structural changes. The BthA-I-BPB structure reveals a novel oligomeric conformation. This conformation is more energetically and conformationally stable than the native structure and the abolition of pharmacological activities by the ligand may be related to the oligomeric structural changes. A residue of the ;pancreatic' loop (Lys69), which is usually attributed as providing the anticoagulant effect, is in the dimeric interface of BthA-I-BPB, leading to a new hypothesis regarding the abolition of this activity by BPB.
- Departamento de Física e Biofísica, Instituto de Biociências, UNESP, Distrito de Rubião Jr s/n, Botucatu-SP, Brazil.
Organizational Affiliation: