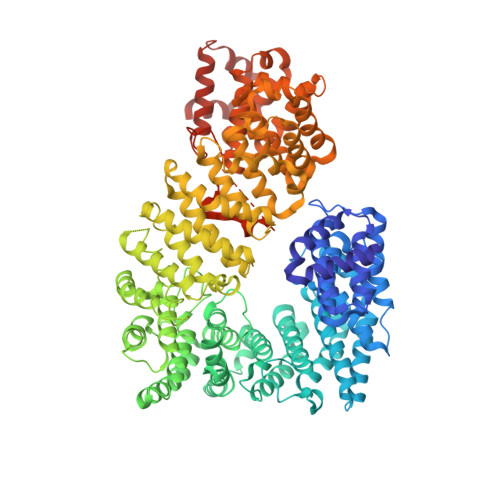
The structure of the nuclear export receptor cse1 in its cytosolic state reveals a closed conformation incompatible with cargo binding
Cook, A., Fernandez, E., Lindner, D., Ebert, J., Schlenstedt, G., Conti, E.(2005) Mol Cell 18: 355-367
- PubMed: 15866177 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.03.021
- Primary Citation Related Structures:
1Z3H - PubMed Abstract:
Cse1 mediates nuclear export of importin alpha, the nuclear localization signal (NLS) import adaptor. We report the 3.1 A resolution structure of cargo-free Cse1, representing this HEAT repeat protein in its cytosolic state. Cse1 is compact, consisting of N- and C-terminal arches that interact to form a ring. Comparison with the structure of cargo-bound Cse1 shows a major conformational change leading to opening of the structure upon cargo binding. The largest structural changes occur within a hinge region centered at HEAT repeat 8. This repeat contains a conserved insertion that connects the RanGTP and importin alpha contact sites and that is essential for binding. In the cargo-free state, the RanGTP binding sites are occluded and the importin alpha sites are distorted. Mutations that destabilize the N- to C-terminal interaction uncouple importin alpha and Ran binding, suggesting that the closed conformation prevents association with importin alpha.
- European Molecular Biology Laboratory, Meyerhofstrasse 1, D-69117 Heidelberg, Germany.
Organizational Affiliation: