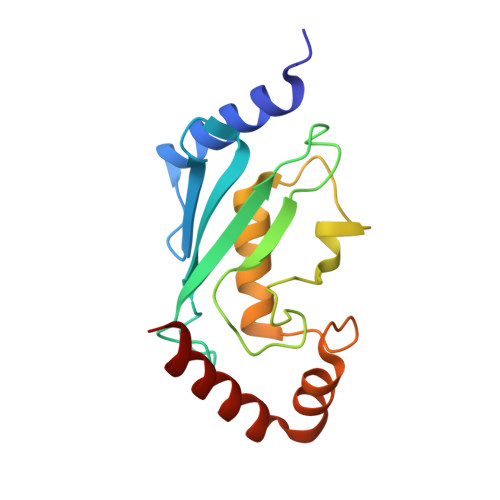
The 1.1A crystallographic structure of ubiquitin-conjugating enzyme (ubc-2) from Caenorhabditis elegans: functional and evolutionary significance
Gavira, J.A., DiGiamamarino, E., Tempel, W., Liu, Z.J., Wang, B.C., Meehan, E., Ng, J.D.To be published.