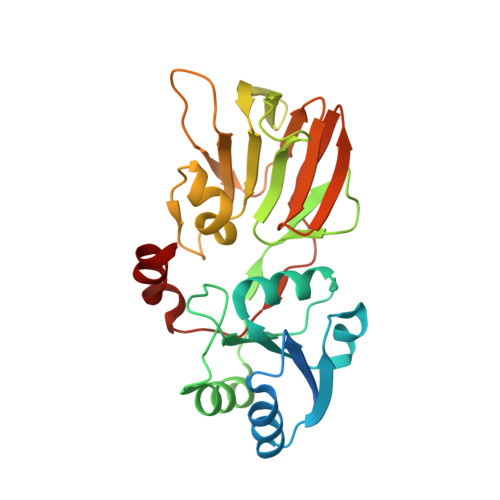
Crystal Structures of an NAD Kinase from Archaeoglobus fulgidus in Complex with ATP, NAD, or NADP
Liu, J., Lou, Y., Yokota, H., Adams, P.D., Kim, R., Kim, S.H.(2005) J Mol Biology 354: 289-303
- PubMed: 16242716 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.09.026
- Primary Citation Related Structures:
1SUW, 1Z0S, 1Z0U, 1Z0Z - PubMed Abstract:
NAD kinase is a ubiquitous enzyme that catalyzes the phosphorylation of NAD to NADP using ATP or inorganic polyphosphate (poly(P)) as phosphate donor, and is regarded as the only enzyme responsible for the synthesis of NADP. We present here the crystal structures of an NAD kinase from the archaeal organism Archaeoglobus fulgidus in complex with its phosphate donor ATP at 1.7 A resolution, with its substrate NAD at 3.05 A resolution, and with the product NADP in two different crystal forms at 2.45 A and 2.0 A resolution, respectively. In the ATP bound structure, the AMP portion of the ATP molecule is found to use the same binding site as the nicotinamide ribose portion of NAD/NADP in the NAD/NADP bound structures. A magnesium ion is found to be coordinated to the phosphate tail of ATP as well as to a pyrophosphate group. The conserved GGDG loop forms hydrogen bonds with the pyrophosphate group in the ATP-bound structure and the 2' phosphate group of the NADP in the NADP-bound structures. A possible phosphate transfer mechanism is proposed on the basis of the structures presented.
- Berkeley Structural Genomics Center, Lawrence Berkeley National Laboratory, Physical Bioscience Division, Berkeley, CA 94720-5230, USA.
Organizational Affiliation: