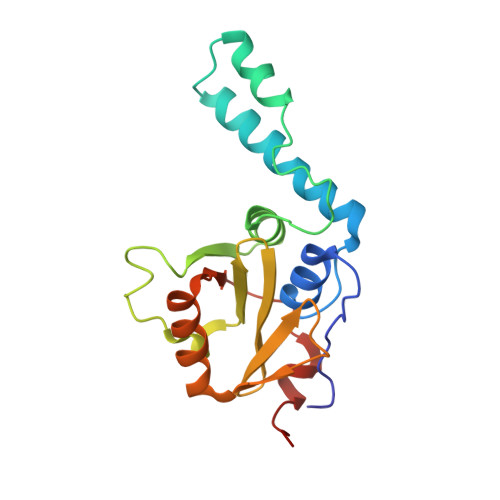
The solution structure of the Josephin domain of ataxin-3: Structural determinants for molecular recognition
Nicastro, G., Menon, R.P., Masino, L., Knowles, P.P., McDonald, N.Q., Pastore, A.(2005) Proc Natl Acad Sci U S A 102: 10493-10498
- PubMed: 16020535 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0501732102
- Primary Citation Related Structures:
1YZB - PubMed Abstract:
The Josephin domain plays an important role in the cellular functions of ataxin-3, the protein responsible for the neurodegenerative Machado-Joseph disease. We have determined the solution structure of Josephin and shown that it belongs to the family of papain-like cysteine proteases, sharing the highest degree of structural similarity with bacterial staphopain. A currently unique structural feature of Josephin is a flexible helical hairpin formed by a 32-residue insertion, which could determine substrate specificity. By using the Josephin structure and the availability of NMR chemical shift assignments, we have mapped the enzyme active site by using the typical cysteine protease inhibitors, transepoxysuccinyl-L-eucylamido-4-guanidino-butane (E-64) and [L-3-trans-(propylcarbamyl)oxirane-2-carbonyl]-L-isoleucyl-L-proline (CA-074). We also demonstrate that the specific interaction of Josephin with the ubiquitin-like domain of the ubiquitin- and proteasome-binding factor HHR23B involves complementary exposed hydrophobic surfaces. The structural similarity with other deubiquitinating enzymes suggests a model for the proteolytic enzymatic activity of ataxin-3.
- National Institute for Medical Research, The Ridgeway, London NW7 1AA, United Kingdom.
Organizational Affiliation: