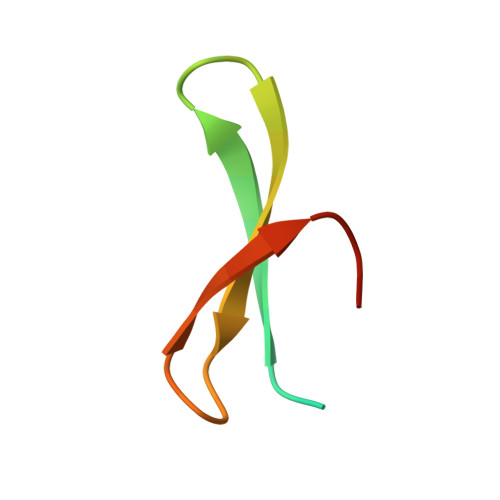
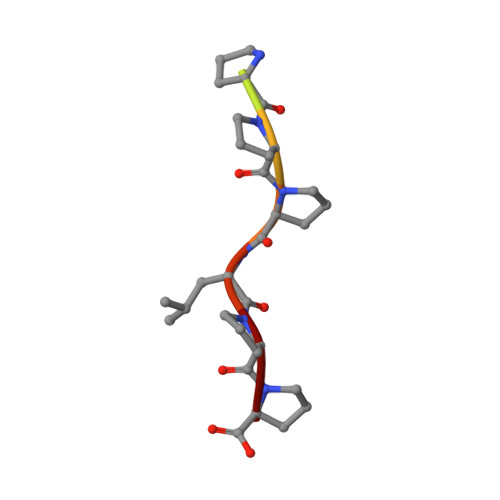
Structural basis for APPTPPPLPP peptide recognition by the FBP11WW1 domain.
Pires, J.R., Parthier, C., Aido-Machado, R., Wiedemann, U., Otte, L., Bohm, G., Rudolph, R., Oschkinat, H.(2005) J Mol Biology 348: 399-408
- PubMed: 15811376 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.02.056
- Primary Citation Related Structures:
1YWI, 1YWJ - PubMed Abstract:
WW domains are small protein-protein interaction modules that recognize proline-rich stretches in proteins. The class II tandem WW domains of the formin binding protein 11 (FBP11) recognize specifically proteins containing PPLPp motifs as present in the formins that are involved in limb and kidney development, and in the methyl-CpG-binding protein 2 (MeCP2), associated with the Rett syndrome. The interaction involves the specific recognition of a leucine side-chain. Here, we report on the novel structure of the complex formed by the FPB11WW1 domain and the formin fragment APPTPPPLPP revealing the specificity determinants of class II WW domains.
- Departamento de Bioquímica Médica, Instituto de Ciências Biomédicas, Universidade Federal do Rio de Janeiro, Av. Brigadeiro Trompowiski s/n CCS, Rio de Janeiro, RJ 21941-590, Brazil. jrmpires@gmx.net
Organizational Affiliation: