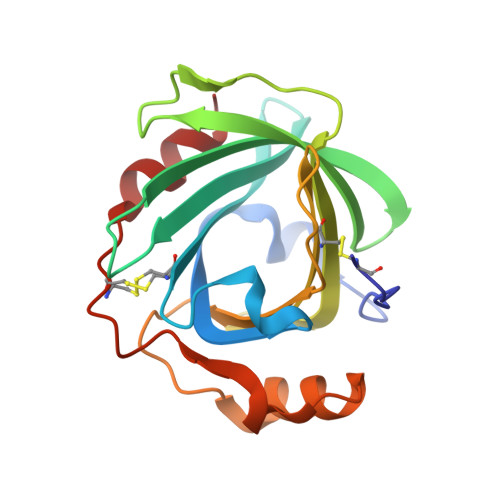
Ultrahigh Resolution Structures of Nitrophorin 4: Heme Distortion in Ferrous CO and NO Complexes
Maes, E.M., Roberts, S.A., Weichsel, A., Montfort, W.R.(2005) Biochemistry 44: 12690-12699
- PubMed: 16171383 Search on PubMed
- DOI: https://doi.org/10.1021/bi0506573
- Primary Citation Related Structures:
1YWA, 1YWB, 1YWC, 1YWD - PubMed Abstract:
Nitrophorin 4 (NP4), a nitric oxide (NO)-transport protein from the blood-sucking insect Rhodnius prolixus, uses a ferric (Fe3+) heme to deliver NO to its victims. NO binding to NP4 induces a large conformational change and complete desolvation of the distal pocket. The heme is markedly nonplanar, displaying a ruffling distortion postulated to contribute to stabilization of the ferric iron. Here, we report the ferrous (Fe2+) complexes of NP4 with NO, CO, and H2O formed after chemical reduction of the protein and the characterization of these complexes by absorption spectroscopy, flash photolysis, and ultrahigh-resolution crystallography (resolutions vary from 0.9 to 1.08 A). The absorption spectra, both in solution and in the crystal, are typical for six-coordinated ferrous complexes. Closure and desolvation of the distal pocket occurs upon binding CO or NO to the iron regardless of the heme oxidation state, confirming that the conformational change is driven by distal ligand polarity. The degree of heme ruffling is coupled to the nature of the ligand and the iron oxidation state in the following order: (Fe3+)-NO > (Fe2+)-NO > (Fe2+)-CO > (Fe3+)-H2O > (Fe2+)-H2O. The ferrous coordination geometry is as expected, except for the proximal histidine bond, which is shorter than typically found in model compounds. These data are consistent with heme ruffling and coordination geometry serving to stabilize the ferric state of the nitrophorins, a requirement for their physiological function. Possible roles for heme distortion and NO bending in heme protein function are discussed.
- Department of Biochemistry and Molecular Biophysics, University of Arizona, Tucson, Arizona 85721, USA.
Organizational Affiliation: