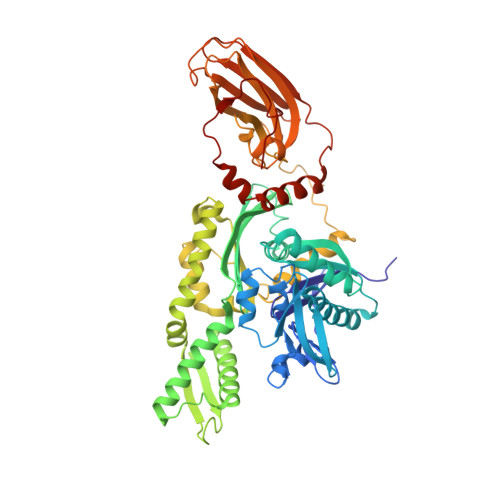
Structural basis of interdomain communication in the Hsc70 chaperone
Jiang, J., Prasad, K., Lafer, E.M., Sousa, R.(2005) Mol Cell 20: 513-524
- PubMed: 16307916 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2005.09.028
- Primary Citation Related Structures:
1YUW - PubMed Abstract:
Hsp70 family proteins are highly conserved chaperones involved in protein folding, degradation, targeting and translocation, and protein complex remodeling. They are comprised of an N-terminal nucleotide binding domain (NBD) and a C-terminal protein substrate binding domain (SBD). ATP binding to the NBD alters SBD conformation and substrate binding kinetics, but an understanding of the mechanism of interdomain communication has been hampered by the lack of a crystal structure of an intact chaperone. We report here the 2.6 angstroms structure of a functionally intact bovine Hsc70 (bHsc70) and a mutational analysis of the observed interdomain interface and the immediately adjacent interdomain linker. This analysis identifies interdomain interactions critical for chaperone function and supports an allosteric mechanism in which the interdomain linker invades and disrupts the interdomain interface when ATP binds.
- Department of Biochemistry, University of Texas Health Science Center, 7703 Floyd Curl Drive, San Antonio, Texas 78229, USA.
Organizational Affiliation: