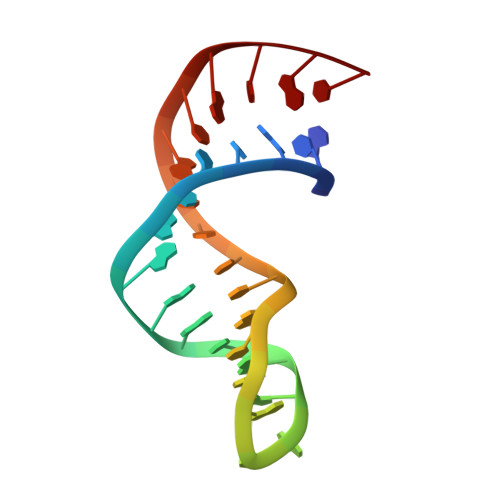
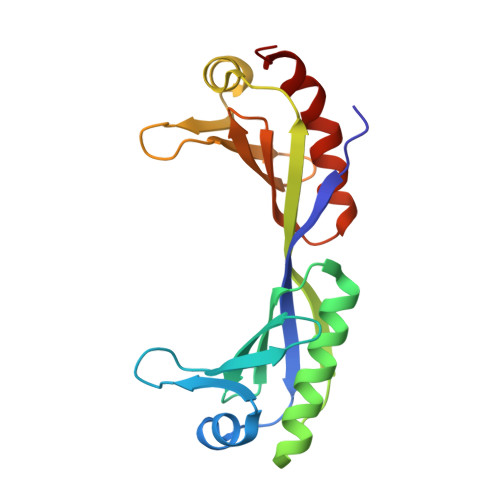
Crystal structure of a yeast TBP/TATA-box complex.
Kim, Y., Geiger, J.H., Hahn, S., Sigler, P.B.(1993) Nature 365: 512-520
- PubMed: 8413604 Search on PubMed
- DOI: https://doi.org/10.1038/365512a0
- Primary Citation Related Structures:
1YTB - PubMed Abstract:
The 2.5 A crystal structure of a TATA-box complex with yeast TBP shows that the eight base pairs of the TATA box bind to the concave surface of TBP by bending towards the major groove with unprecedented severity. This produces a wide open, underwound, shallow minor groove which forms a primarily hydrophobic interface with the entire under-surface of the TBP saddle. The severe bend and a positive writhe radically alter the trajectory of the flanking B-form DNA.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, Connecticut 06510.
Organizational Affiliation: