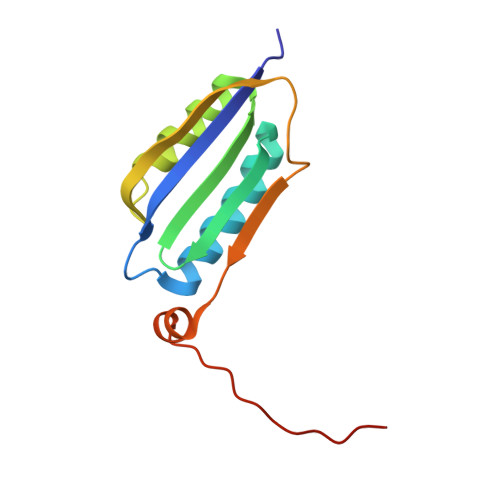
Structure of a Novel Photoreceptor, the BLUF Domain of AppA from Rhodobacter sphaeroides
Anderson, S., Dragnea, V., Masuda, S., Ybe, J., Moffat, K., Bauer, C.(2005) Biochemistry 44: 7998-8005
- PubMed: 15924418 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi0502691
- Primary Citation Related Structures:
1YRX - PubMed Abstract:
The flavin-binding BLUF domain of AppA represents a new class of blue light photoreceptors that are present in a number of bacterial and algal species. The dark state X-ray structure of this domain was determined at 2.3 A resolution. The domain demonstrates a new function for the common ferredoxin-like fold; two long alpha-helices flank the flavin, which is bound with its isoalloxazine ring perpendicular to a five-stranded beta-sheet. The hydrogen bond network and the overall protein topology of the BLUF domain (but not its sequence) bear some resemblance to LOV domains, a subset of PAS domains widely involved in signaling. Nearly all residues conserved in BLUF domains surround the flavin chromophore, many of which are involved in an intricate hydrogen bond network. Photoactivation may induce a rearrangement in this network via reorientation of the Gln63 side chain to form a new hydrogen bond to the flavin O4 position. This shift would also break a hydrogen bond to the Trp104 side chain, which may be critical in induction of global structural change in AppA.
- Consortium for Advanced Radiation Sources, Department of Biochemistry and Molecular Biology, University of Chicago, Chicago, Illinois 60637, USA. smander@uchicago.edu
Organizational Affiliation: