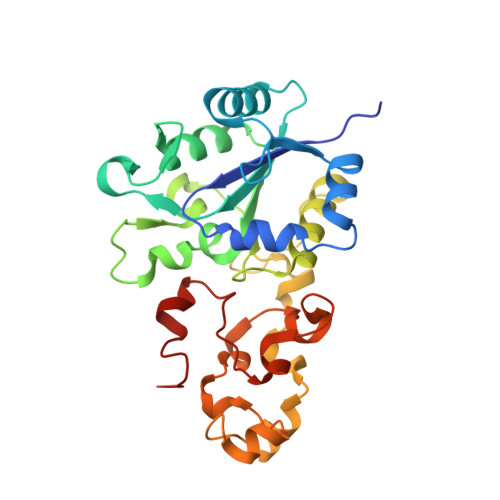
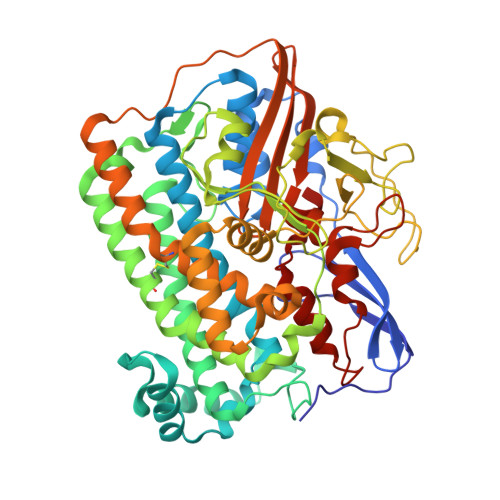
Structural differences between the ready and unready oxidized states of [NiFe] hydrogenases.
Volbeda, A., Martin, L., Cavazza, C., Matho, M., Faber, B.W., Roseboom, W., Albracht, S.P., Garcin, E., Rousset, M., Fontecilla-Camps, J.C.(2005) J Biol Inorg Chem 10: 239-249
- PubMed: 15803334 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-005-0632-x
- Primary Citation Related Structures:
1YQ9, 1YQW, 1YRQ - PubMed Abstract:
[NiFe] hydrogenases catalyze the reversible heterolytic cleavage of molecular hydrogen. Several oxidized, inactive states of these enzymes are known that are distinguishable by their very different activation properties. So far, the structural basis for this difference has not been understood because of lack of relevant crystallographic data. Here, we present the crystal structure of the ready Ni-B state of Desulfovibrio fructosovorans [NiFe] hydrogenase and show it to have a putative mu-hydroxo Ni-Fe bridging ligand at the active site. On the other hand, a new, improved refinement procedure of the X-ray diffraction data obtained for putative unready Ni-A/Ni-SU states resulted in a more elongated electron density for the bridging ligand, suggesting that it is a diatomic species. The slow activation of the Ni-A state, compared with the rapid activation of the Ni-B state, is therefore proposed to result from the different chemical nature of the ligands in the two oxidized species. Our results along with very recent electrochemical studies suggest that the diatomic ligand could be hydro-peroxide.
- Laboratoire de Cristallographie et de Cristallogenèse des Protèines, Institut de Biologie Structurale J.P. Ebel (CEA-CNRS-UJF), 41 rue Jules Horowitz, 38027, Grenoble Cédex 1, France. anne.volbeda@ibs.fr
Organizational Affiliation: