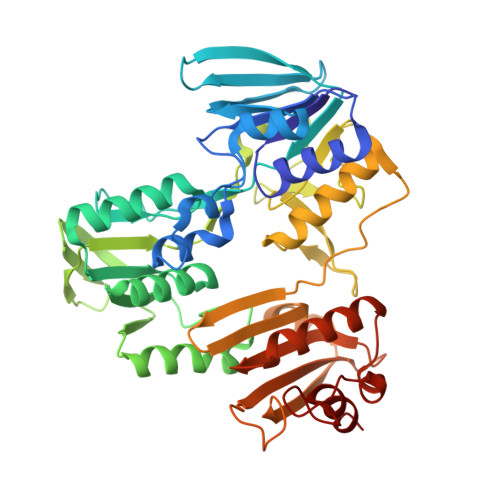
Structure of coenzyme A-disulfide reductase from Staphylococcus aureus at 1.54 A resolution.
Mallett, T.C., Wallen, J.R., Karplus, P.A., Sakai, H., Tsukihara, T., Claiborne, A.(2006) Biochemistry 45: 11278-11289
- PubMed: 16981688 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi061139a
- Primary Citation Related Structures:
1YQZ - PubMed Abstract:
Coenzyme A (CoASH) replaces glutathione as the major low molecular weight thiol in Staphylococcus aureus; it is maintained in the reduced state by coenzyme A-disulfide reductase (CoADR), a homodimeric enzyme similar to NADH peroxidase but containing a novel Cys43-SSCoA redox center. The crystal structure of S. aureus CoADR has been solved using multiwavelength anomalous dispersion data and refined at a resolution of 1.54 A. The resulting electron density maps define the Cys43-SSCoA disulfide conformation, with Cys43-S(gamma) located at the flavin si face, 3.2 A from FAD-C4aF, and the CoAS- moiety lying in an extended conformation within a cleft at the dimer interface. A well-ordered chloride ion is positioned adjacent to the Cys43-SSCoA disulfide and receives a hydrogen bond from Tyr361'-OH of the complementary subunit, suggesting a role for Tyr361' as an acid-base catalyst during the reduction of CoAS-disulfide. Tyr419'-OH is located 3.2 A from Tyr361'-OH as well and, based on its conservation in known functional CoADRs, also appears to be important for activity. Identification of residues involved in recognition of the CoAS-disulfide substrate and in formation and stabilization of the Cys43-SSCoA redox center has allowed development of a CoAS-binding motif. Bioinformatics analyses indicate that CoADR enzymes are broadly distributed in both bacterial and archaeal kingdoms, suggesting an even broader significance for the CoASH/CoAS-disulfide redox system in prokaryotic thiol/disulfide homeostasis.
- Center for Structural Biology, Wake Forest University School of Medicine, Winston-Salem, North Carolina 27157, USA.
Organizational Affiliation: