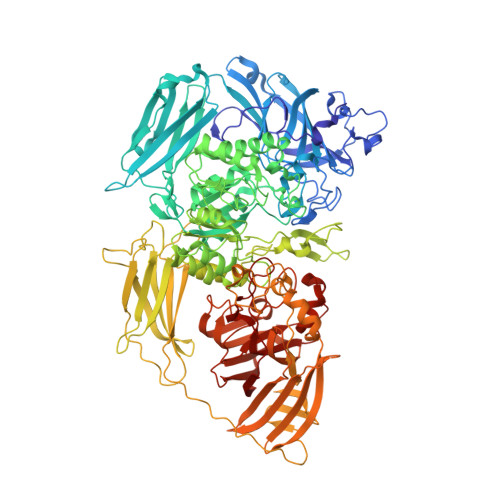
Cold-active beta-Galactosidase from Arthrobacter sp. C2-2 Forms Compact 660kDa Hexamers: Crystal Structure at 1.9A Resolution
Skalova, T., Dohnalek, J., Spiwok, V., Lipovova, P., Vondrackova, E., Petrokova, H., Duskova, J., Strnad, H., Kralova, B., Hasek, J.(2005) J Mol Biology 353: 282-294
- PubMed: 16171818 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.08.028
- Primary Citation Related Structures:
1YQ2 - PubMed Abstract:
The X-ray structure of cold-active beta-galactosidase (isoenzyme C-2-2-1) from an Antarctic bacterium Arthrobacter sp. C2-2 was solved at 1.9A resolution. The enzyme forms 660 kDa hexamers with active sites opened to the central cavity of the hexamer and connected by eight channels with exterior solvent. To our best knowledge, this is the first cold-active beta-galactosidase with known structure and also the first known beta-galactosidase structure in the form of compact hexamers. The hexamer organization regulates access of substrates and ligands to six active sites and this unique packing, present also in solution, raises questions about its purpose and function. This enzyme belongs to glycosyl hydrolase family 2, similarly to Escherichia coli beta-galactosidase, forming tetramers necessary for its enzymatic function. However, we discovered significant differences between these two enzymes affecting the ability of tetramer/hexamer formation and complementation of the active site. This structure reveals new insights into the cold-adaptation mechanisms of enzymatic pathways of extremophiles.
- Institute of Macromolecular Chemistry, Academy of Sciences of the Czech Republic, Heyrovského nám. 2, 1606 Praha 6, Czech Republic. skalova@imc.cas.cz
Organizational Affiliation: