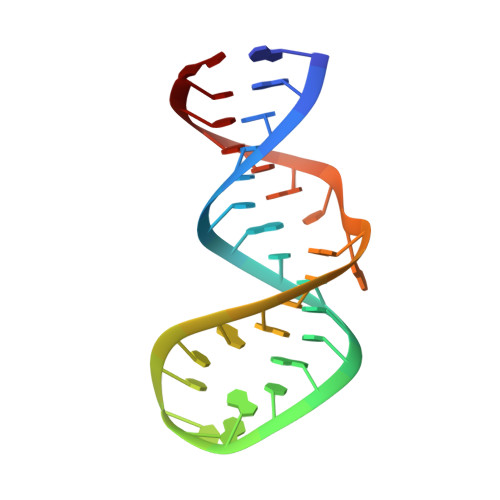
NMR structure of the apoB mRNA stem-loop and its interaction with the C to U editing APOBEC1 complementary factor.
Maris, C., Masse, J., Chester, A., Navaratnam, N., Allain, F.H.(2005) RNA 11: 173-186
- PubMed: 15659357 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1261/rna.7190705
- Primary Citation Related Structures:
1YLG, 1YNC, 1YNE, 1YNG - PubMed Abstract:
We have solved the NMR structure of the 31-nucleotide (nt) apoB mRNA stem-loop, a substrate of the cytidine deaminase APOBEC1. We found that the edited base located at the 5' end of the octa-loop is stacked between two adenosines in both the unedited (cytidine 6666) and the edited (uridine 6666) forms and that the rest of the loop is unstructured. The 11-nt "mooring" sequence essential for editing is partially flexible although it is mostly in the stem of the RNA. The octa-loop and the internal loop in the middle of the stem confer this flexibility. These findings shed light on why APOBEC1 alone cannot edit efficiently the cytidine 6666 under physiological conditions, the editing base being buried in the loop and not directly accessible. We also show that APOBEC1 does not specifically bind apoB mRNA and requires the auxiliary factor, APOBEC1 complementary factor (ACF), to edit specifically cytidine 6666. The binding of ACF to both the mooring sequence and APOBEC1 explains the specificity of the reaction. Our NMR study lead us to propose a mechanism in which ACF recognizes first the flexible nucleotides of the mooring sequence (the internal loop and the 3' end octa-loop) and subsequently melts the stem-loop, exposing the amino group of the cytidine 6666 to APOBEC1. Thus, the flexibility of the mooring sequence plays a central role in the RNA recognition by ACF.
- Institute for Molecular Biology and Biophysics, ETH Hönggerberg HPK D11.2, CH-8093 Zürich, Switzerland.
Organizational Affiliation: