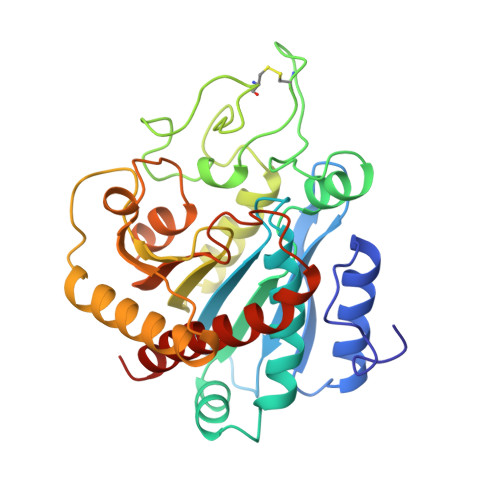
Carboxypeptidase A: native, zinc-removed and mercury-replaced forms.
Greenblatt, H.M., Feinberg, H., Tucker, P.A., Shoham, G.(1998) Acta Crystallogr D Biol Crystallogr 54: 289-305
- PubMed: 9867434 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444997010445
- Primary Citation Related Structures:
1ARL, 1ARM, 1YME - PubMed Abstract:
The crystal structure of the zinc-containing exopeptidase bovine carboxypeptidase A (CPA) has been refined to high resolution, based on a data set collected from a single crystal, incorporating new sequence information based on cloning of the bovine gene. In addition, new refined structures are available for the zinc-removed form of the enzyme, apo-CPA, as well as the mercury-replaced form, Hg-CPA. The native structure reveals that the zinc-bound water molecule does not appear to more loosely bound than the rest of the zinc ligands, at least when B-factor values are considered. Nor is there any evidence for a secondary location of this water molecule. The apo-enzyme structure does not show any density in the place of the removed zinc ion. The only significant change appears to be a chi2 rotation of one zinc histidine ligand to form an ion-pair interaction with a glutamic acid side chain. The structure of Hg-CPA reveals a solvent Tris molecule bound to the mercury cation, as well as an unidentified cation bound to Glu270. The location of this citation agrees with previous proposals for the binding side of inhibitory zinc. These observations may explain some of the differences in kinetics observed in metal- replaced CPA.
- Department of Inorganic and Analytical Chemistry, Hebrew University, Jerusalem, Israel, 91904.
Organizational Affiliation: