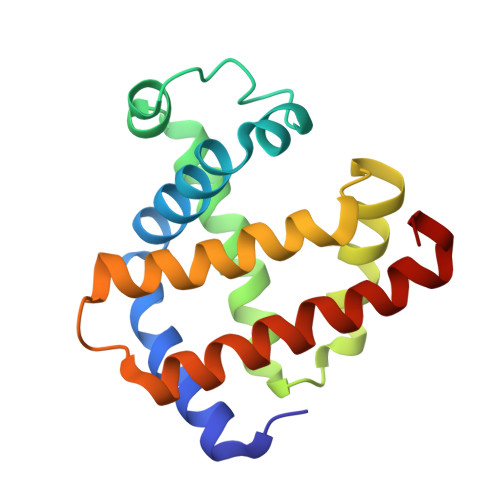
Three-dimensional structure of cyanomet-sulfmyoglobin C.
Evans, S.V., Sishta, B.P., Mauk, A.G., Brayer, G.D.(1994) Proc Natl Acad Sci U S A 91: 4723-4726
- PubMed: 8197124 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.91.11.4723
- Primary Citation Related Structures:
1YMC - PubMed Abstract:
The atomic structure of horse heart cyanomet-sulfmyoglobin C has been established by x-ray crystallographic techniques to a resolution of 2.0 A with an R value of 0.129. The protoheme IX prosthetic group of this thermodynamically stable sulfmyoglobin derivative has been converted to a chlorin in which the pyrrole ring bearing the 4-vinyl group is saturated and possesses an exocyclic thiolene ring. This study provides the three-dimensional structure of a protein with an iron-chlorin prosthetic group. The overall conformation of the surrounding polypeptide chain of the modified protein is very similar to that of the native protein. However, the addition of the sulfur atom has caused a distortion of the prosthetic group from that in the native protein to result in the repositioning of the side chains of some residues in the heme pocket.
- Department of Biochemistry and Molecular Biology, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: