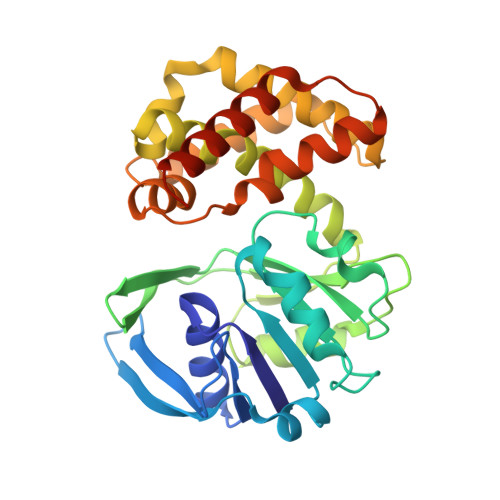
The crystal structure of Escherichia coli ketopantoate reductase with NADP+ bound.
Lobley, C.M., Ciulli, A., Whitney, H.M., Williams, G., Smith, A.G., Abell, C., Blundell, T.L.(2005) Biochemistry 44: 8930-8939
- PubMed: 15966718 Search on PubMed
- DOI: https://doi.org/10.1021/bi0502036
- Primary Citation Related Structures:
1YJQ - PubMed Abstract:
The NADPH-dependent reduction of ketopantoate to pantoate, catalyzed by ketopantoate reductase (KPR; EC 1.1.1.169), is essential for the biosynthesis of pantothenate (vitamin B(5)). Here we present the crystal structure of Escherichia coli KPR with NADP(+) bound, solved to 2.1 A resolution. The cofactor is bound in the active site cleft between the N-terminal Rossmann-fold domain and the C-terminal alpha-helical domain. The thermodynamics of cofactor and substrate binding were characterized by isothermal titration calorimetry. The dissociation constant for NADP(+) was found to be 6.5 muM, 20-fold larger than that for NADPH (0.34 muM). The difference is primarily due to the entropic term, suggesting favorable hydrophobic interactions of the more lipophilic nicotinamide ring in NADPH. Comparison of this binary complex structure with the previously studied apoenzyme reveals no evidence for large domain movements on cofactor binding. This observation is further supported both by molecular dynamics and by calorimetric analysis. A model of the ternary complex, based on the structure presented here, provides novel insights into the molecular mechanism of enzyme catalysis. We propose a conformational switch of the essential Lys176 from the "resting" state observed in our structure to an "active" state, to bind ketopantoate. Additionally, we identify the importance of Asn98 for substrate binding and enzyme catalysis.
- Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge CB2 1GA, UK.
Organizational Affiliation: