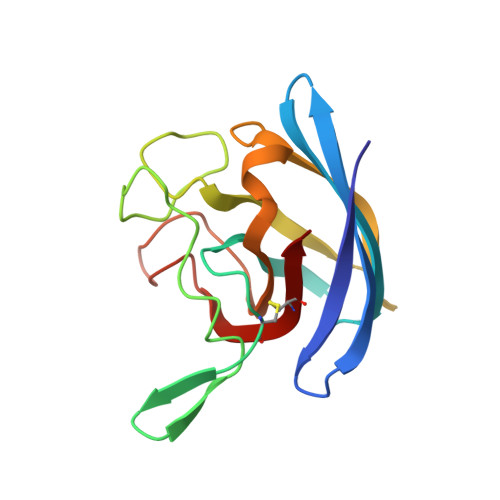
Novel dimeric interface and electrostatic recognition in bacterial Cu,Zn superoxide dismutase.
Bourne, Y., Redford, S.M., Steinman, H.M., Lepock, J.R., Tainer, J.A., Getzoff, E.D.(1996) Proc Natl Acad Sci U S A 93: 12774-12779
- PubMed: 8917495 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.93.23.12774
- Primary Citation Related Structures:
1YAI - PubMed Abstract:
Eukaryotic Cu,Zn superoxide dismutases (CuZnSODs) are antioxidant enzymes remarkable for their unusually stable beta-barrel fold and dimer assembly, diffusion-limited catalysis, and electrostatic guidance of their free radical substrate. Point mutations of CuZnSOD cause the fatal human neurodegenerative disease amyotrophic lateral sclerosis. We determined and analyzed the first crystallographic structure (to our knowledge) for CuZnSOD from a prokaryote, Photobacterium leiognathi, a luminescent symbiont of Leiognathid fish. This structure, exemplifying prokaryotic CuZnSODs, shares the active-site ligand geometry and the topology of the Greek key beta-barrel common to the eukaryotic CuZnSODs. However, the beta-barrel elements recruited to form the dimer interface, the strategy used to forge the channel for electrostatic recognition of superoxide radical, and the connectivity of the intrasubunit disulfide bond in P. leiognathi CuZnSOD are discrete and strikingly dissimilar from those highly conserved in eukaryotic CuZnSODs. This new CuZnSOD structure broadens our understanding of structural features necessary and sufficient for CuZnSOD activity, highlights a hitherto unrecognized adaptability of the Greek key beta-barrel building block in evolution, and reveals that prokaryotic and eukaryotic enzymes diverged from one primordial CuZnSOD and then converged to distinct dimeric enzymes with electrostatic substrate guidance.
- Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: