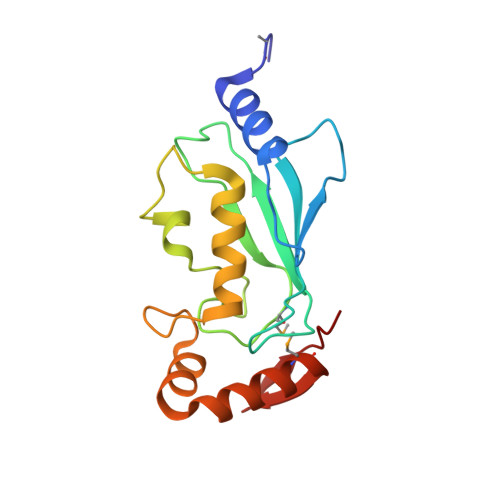
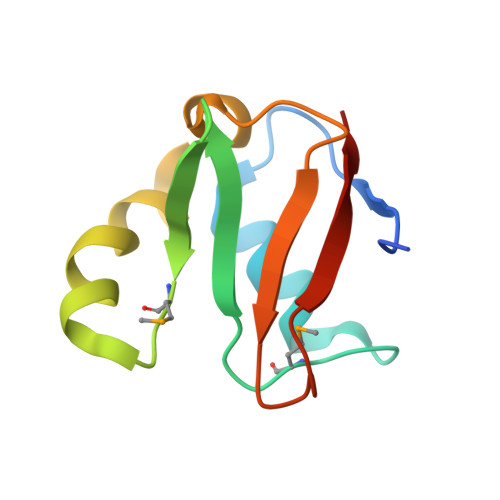
Structural basis for recruitment of Ubc12 by an E2 binding domain in NEDD8's E1.
Huang, D.T., Paydar, A., Zhuang, M., Waddell, M.B., Holton, J.M., Schulman, B.A.(2005) Mol Cell 17: 341-350
- PubMed: 15694336
- DOI: https://doi.org/10.1016/j.molcel.2004.12.020
- Primary Citation of Related Structures:
1Y8X - PubMed Abstract:
E2 conjugating enzymes play a central role in ubiquitin and ubiquitin-like protein (ublp) transfer cascades: the E2 accepts the ublp from the E1 enzyme and then the E2 often interacts with an E3 enzyme to promote ublp transfer to the target. We report here the crystal structure of a complex between the C-terminal domain from NEDD8's heterodimeric E1 (APPBP1-UBA3) and the catalytic core domain of NEDD8's E2 (Ubc12). The structure and associated mutational analyses reveal molecular details of Ubc12 recruitment by NEDD8's E1. Interestingly, the E1's Ubc12 binding domain resembles ubiquitin and recruits Ubc12 in a manner mimicking ubiquitin's interactions with ubiquitin binding domains. Structural comparison with E2-E3 complexes indicates that the E1 and E3 binding sites on Ubc12 may overlap and raises the possibility that crosstalk between E1 and E3 interacting with an E2 could influence the specificity and processivity of ublp transfer.
- Department of Structural Biology, Department of Genetics/Tumor Cell Biology, St. Jude Children's Research Hospital, Memphis, TN 38105, USA.
Organizational Affiliation: