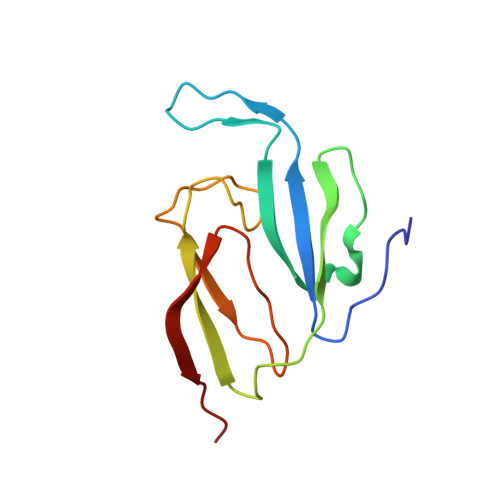
Solution structure of a two-repeat fragment of major vault protein.
Kozlov, G., Vavelyuk, O., Minailiuc, O., Banville, D., Gehring, K., Ekiel, I.(2006) J Mol Biology 356: 444-452
- PubMed: 16373071 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.11.064
- Primary Citation Related Structures:
1Y7X - PubMed Abstract:
Major vault protein (MVP) is the main constituent of vaults, large ribonucleoprotein particles implicated in resistance to cancer therapy and correlated with poor survival prognosis. Here, we report the structure of the main repeat element in human MVP. The approximately 55 amino acid residue MVP domain has a unique, novel fold that consists of a three-stranded antiparallel beta-sheet. The solution NMR structure of a two-domain fragment reveals the interdomain contacts and relative orientations of the two MVP domains. We use these results to model the assembly of 672 MVP domains from 96 MVP molecules into the ribs of the 13MDa vault structure. The unique features include a thin, skin-like structure with polar residues on both the cytoplasmic and internal surface, and a pole-to-pole arrangement of MVP molecules. These studies provide a starting point for understanding the self-assembly of MVP into vaults and their interactions with other proteins. Chemical shift perturbation studies identified the binding site of vault poly(ADP-ribose) polymerase, another component of vault particles, indicating that MVP domains form a new class of interaction-mediating modules.
- Department of Biochemistry, McGill University, 3655 Promenade Sir William Osler, Montréal, Que., Canada.
Organizational Affiliation: