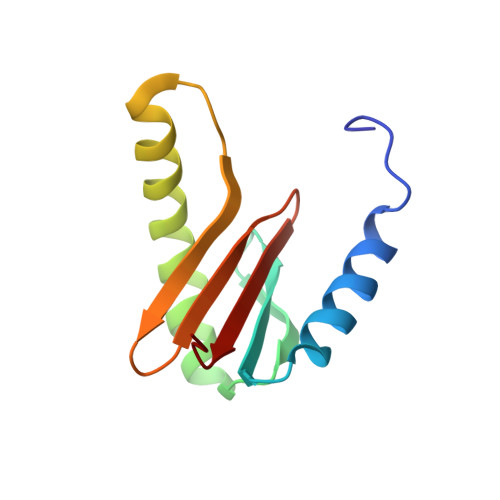
Solution structure of isoform 1 of Roadblock/LC7, a light chain in the dynein complex.
Song, J., Tyler, R.C., Lee, M.S., Tyler, E.M., Markley, J.L.(2005) J Mol Biology 354: 1043-1051
- PubMed: 16289575 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.10.017
- Primary Citation Related Structures:
1Y4O - PubMed Abstract:
Roadblock/LC7 is a member of a class of dynein light chains involved in regulating the function of the dynein complex. We have determined the three-dimensional structure of isoform 1 of the mouse Roadblock/LC7 cytoplasmic dynein light chain (robl1_mouse) by NMR spectroscopy. In contrast to a previously reported NMR structure of the human homolog with 96% sequence identity (PDB 1TGQ), which showed the protein as a monomer, our results indicate clearly that robl1 exists as a symmetric homodimer. The two beta3-strands pair with each other and form a continuous ten-stranded beta-sheet. The 25-residue alpha2-helix from one subunit packs antiparallel to that of the other subunit on the face of the beta-sheet. Zipper-like hydrophobic contacts between the two helices serve to stabilize the dimer. Through an NMR titration experiment, we localized the site on robl1_mouse that interacts with the 40 residue peptide spanning residues 243 through 282 of IC74-1_rat. These results provide physical evidence for a symmetrical interaction between dimeric robl1 and the two molecules of IC74-1 in the dynein complex.
- Center for Eukaryotic Structural Genomics, Department of Biochemistry, University of Wisconsin-Madison, WI 53706-1544, USA.
Organizational Affiliation: