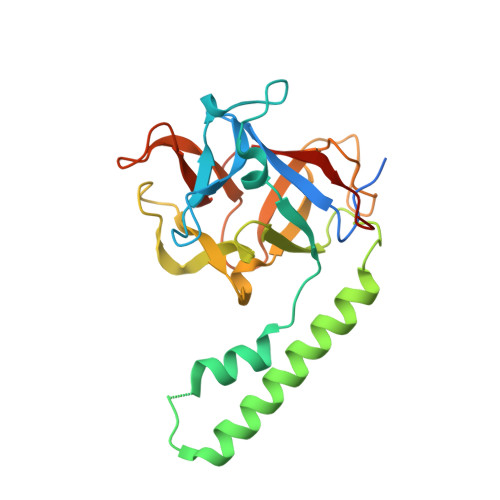
Crystal structure of the ligand binding suppressor domain of type 1 inositol 1,4,5-trisphosphate receptor.
Bosanac, I., Yamazaki, H., Matsu-Ura, T., Michikawa, T., Mikoshiba, K., Ikura, M.(2005) Mol Cell 17: 193-203
- PubMed: 15664189 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2004.11.047
- Primary Citation Related Structures:
1XZZ - PubMed Abstract:
Binding of inositol 1,4,5-trisphosphate (IP(3)) to the amino-terminal region of IP(3) receptor promotes Ca(2+) release from the endoplasmic reticulum. Within the amino terminus, the first 220 residues directly preceding the IP(3) binding core domain play a key role in IP(3) binding suppression and regulatory protein interaction. Here we present a crystal structure of the suppressor domain of the mouse type 1 IP(3) receptor at 1.8 A. Displaying a shape akin to a hammer, the suppressor region contains a Head subdomain forming the beta-trefoil fold and an Arm subdomain possessing a helix-turn-helix structure. The conserved region on the Head subdomain appeared to interact with the IP(3) binding core domain and is in close proximity to the previously proposed binding sites of Homer, RACK1, calmodulin, and CaBP1. The present study sheds light onto the mechanism underlying the receptor's sensitivity to the ligand and its communication with cellular signaling proteins.
- Division of Molecular and Structural Biology, Ontario Cancer Institute and Department of Medical Biophysics, University of Toronto, 610 University Avenue, Toronto, Ontario, M5G 2M9, Canada.
Organizational Affiliation: