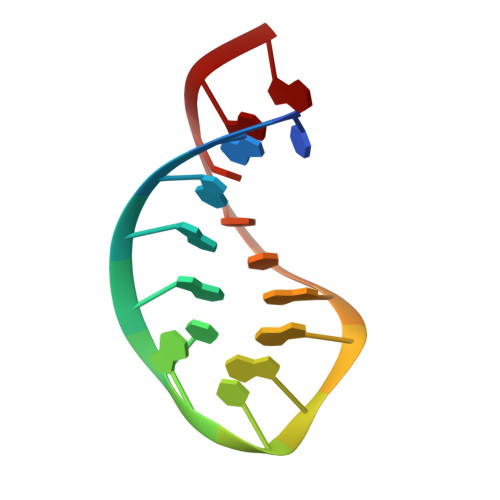
NMR structures of double loops of an RNA aptamer against mammalian initiation factor 4A
Sakamoto, T., Oguro, A., Kawai, G., Ohtsu, T., Nakamura, Y.(2005) Nucleic Acids Res 33: 745-754
- PubMed: 15687383 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gki222
- Primary Citation Related Structures:
1XWP, 1XWU - PubMed Abstract:
A high affinity RNA aptamer (APT58, 58 nt long) against mammalian initiation factor 4A (eIF4A) requires nearly its entire nucleotide sequence for efficient binding. Since splitting either APT58 or eIF4A into two domains diminishes the affinity for each other, it is suggested that multiple interactions or a global interaction between the two molecules accounts for the high affinity. To understand the structural basis of APT58's global recognition of eIF4A, we determined the solution structure of two essential nucleotide loops (AUCGCA and ACAUAGA) within the aptamer using NMR spectroscopy. The AUCGCA loop is stabilized by a U-turn motif and contains a non-canonical A:A base pair (the single hydrogen bond mismatch: Hoogsteen/Sugar-edge). On the other hand, the ACAUAGA loop is stabilized by an AUA tri-nucleotide loop motif and contains the other type of A:A base pair (single hydrogen bond mismatch: Watson-Crick/Watson-Crick). Considering the known structural and functional properties of APT58, we propose that the AUCGCA loop is directly involved in the interaction with eIF4A, while the flexibility of the ACAUAGA loop is important to support this interaction. The Watson-Crick edges of C7 and C9 in the AUCGCA loop may directly interact with eIF4A.
- Department of Basic Medical Sciences, Institute of Medical Science, University of Tokyo 4-6-1 Shirokanedai, Minato-ku, Tokyo 108-8639, Japan.
Organizational Affiliation: