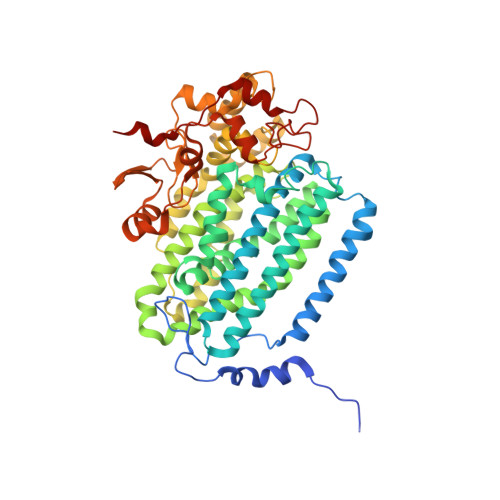
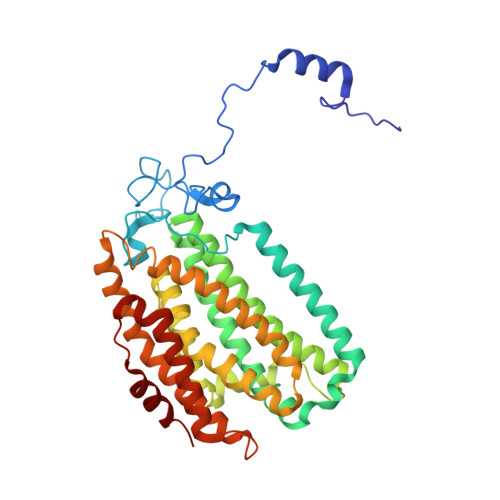
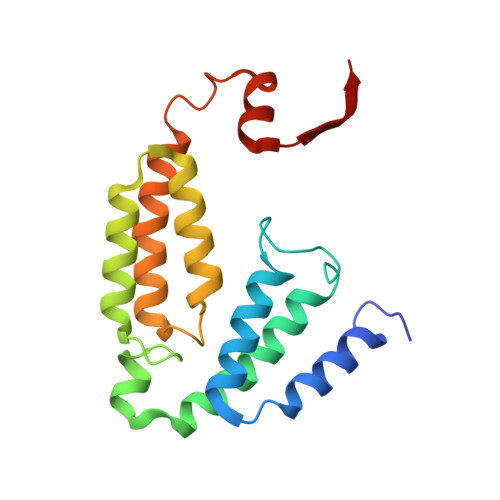
Product Bound Structures of the Soluble Methane Monooxygenase Hydroxylase from Methylococcus capsulatus (Bath): Protein Motion in the Alpha-Subunit
Sazinsky, M.H., Lippard, S.J.(2005) J Am Chem Soc 127: 5814-5825
- PubMed: 15839679 Search on PubMed
- DOI: https://doi.org/10.1021/ja044099b
- Primary Citation Related Structures:
1XU3, 1XU5, 1XVB, 1XVC, 1XVD, 1XVE, 1XVF, 1XVG - PubMed Abstract:
The soluble methane monooxygenase hydroxylase (MMOH) alpha-subunit contains a series of cavities that delineate the route of substrate entrance to and product egress from the buried carboxylate-bridged diiron center. The presence of discrete cavities is a major structural difference between MMOH, which can hydroxylate methane, and toluene/o-xylene monooxygenase hydroxylase (ToMOH), which cannot. To understand better the functions of the cavities and to investigate how an enzyme designed for methane hydroxylation can also accommodate larger substrates such as octane, methylcubane, and trans-1-methyl-2-phenylcyclopropane, MMOH crystals were soaked with an assortment of different alcohols and their X-ray structures were solved to 1.8-2.4 A resolution. The product analogues localize to cavities 1-3 and delineate a path of product exit and/or substrate entrance from the active site to the surface of the protein. The binding of the alcohols to a position bridging the two iron atoms in cavity 1 extends and validates previous crystallographic, spectroscopic, and computational work indicating this site to be where substrates are hydroxylated and products form. The presence of these alcohols induces perturbations in the amino acid side-chain gates linking pairs of cavities, allowing for the formation of a channel similar to one observed in ToMOH. Upon binding of 6-bromohexan-1-ol, the pi helix formed by residues 202-211 in helix E of the alpha-subunit is extended through residue 216, changing the orientations of several amino acid residues in the active site cavity. This remarkable secondary structure rearrangement in the four-helix bundle has several mechanistic implications for substrate accommodation and the function of the effector protein, MMOB.
- Contribution from the Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: