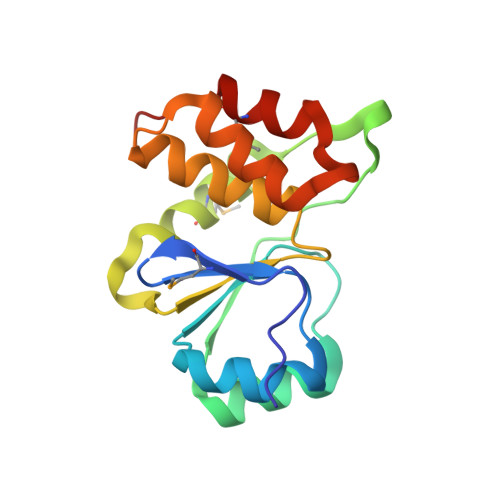
Structural and functional characterization of a novel phosphatase from the Arabidopsis thaliana gene locus At1g05000.
Aceti, D.J., Bitto, E., Yakunin, A.F., Proudfoot, M., Bingman, C.A., Frederick, R.O., Sreenath, H.K., Vojtik, F.C., Wrobel, R.L., Fox, B.G., Markley, J.L., Phillips Jr., G.N.(2008) Proteins 73: 241-253
- PubMed: 18433060 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/prot.22041
- Primary Citation Related Structures:
1XRI - PubMed Abstract:
The crystal structure of the protein product of the gene locus At1g05000, a hypothetical protein from A. thaliana, was determined by the multiple-wavelength anomalous diffraction method and was refined to an R factor of 20.4% (R(free) = 24.9%) at 3.3 A. The protein adopts the alpha/beta fold found in cysteine phosphatases, a superfamily of phosphatases that possess a catalytic cysteine and form a covalent thiol-phosphate intermediate during the catalytic cycle. In At1g05000, the analogous cysteine (Cys(150)) is located at the bottom of a positively-charged pocket formed by residues that include the conserved arginine (Arg(156)) of the signature active site motif, HCxxGxxRT. Of 74 model phosphatase substrates tested, purified recombinant At1g05000 showed highest activity toward polyphosphate (poly-P(12-13)) and deoxyribo- and ribonucleoside triphosphates, and less activity toward phosphoenolpyruvate, phosphotyrosine, phosphotyrosine-containing peptides, and phosphatidyl inositols. Divalent metal cations were not required for activity and had little effect on the reaction.
- Department of Biochemistry, The Center for Eukaryotic Structural Genomics, University of Wisconsin at Madison, Madison, Wisconsin 53706, USA.
Organizational Affiliation: