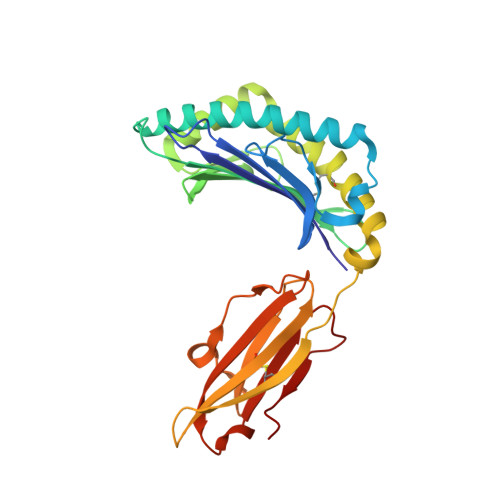
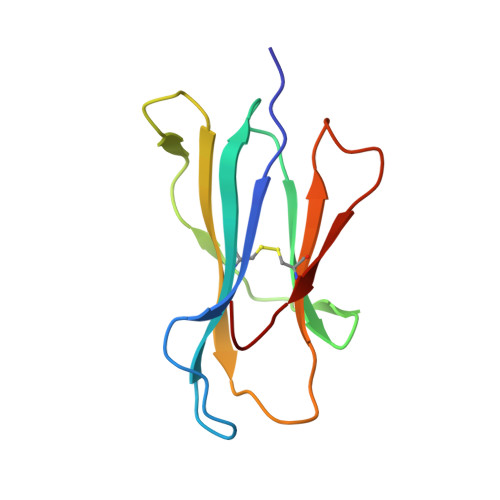
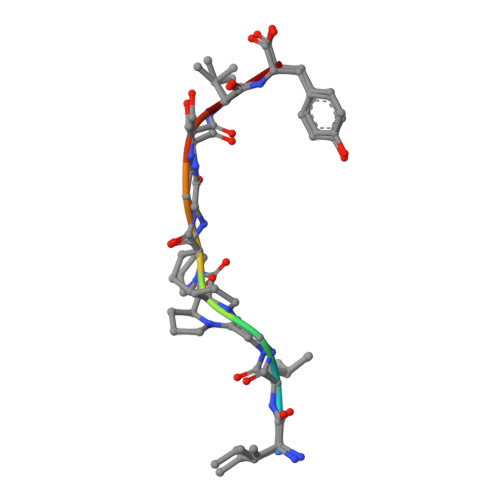
Crystal structures of two peptide-HLA-B*1501 complexes; structural characterization of the HLA-B62 supertype
Roder, G., Blicher, T., Justesen, S., Johannesen, B., Kristensen, O., Kastrup, J., Buus, S., Gajhede, M.(2006) Acta Crystallogr D Biol Crystallogr 62: 1300-1310
- PubMed: 17057332
- DOI: https://doi.org/10.1107/S0907444906027636
- Primary Citation Related Structures:
1XR8, 1XR9 - PubMed Abstract:
MHC class I molecules govern human cytotoxic T cell responses. Their specificity determines which peptides they sample from the intracellular protein environment and then present to human cytotoxic T cells. More than 1100 different MHC class I proteins have been found in human populations and it would be a major undertaking to address each of these specificities individually. Based upon their peptide binding specificity, they are currently subdivided into 12 supertypes. Several of these HLA supertypes have not yet been described at the structural level. To support a comprehensive understanding of human immune responses, the structure of at least one member of each supertype should be determined. Here, the structures of two immunogenic peptide-HLA-B*1501 complexes are described. The structure of HLA-B*1501 in complex with a peptide (LEKARGSTY, corresponding to positions 274-282 in the Epstein-Barr virus nuclear antigen-3A) was determined to 2.3 A resolution. The structure of HLA-B*1501 in complex with a peptide (ILGPPGSVY) derived from human ubiquitin-conjugating enzyme-E2 corresponding to positions 91-99 was solved to 1.8 A resolution. Mutual comparisons of these two structures with structures from other HLA supertypes define and explain the specificity of the P2 and P9 peptide anchor preferences in the B62 HLA supertype. The P2 peptide residue binds to the B-pocket in HLA-B*1501. This pocket is relatively large because of the small Ser67 residue located at the bottom. The peptide proximal part of the B-pocket is hydrophobic, which is consistent with P2 anchor residue preference for Leu. The specificity of the B-pocket is determined by the Met45, Ile66 and Ser67 residues. The apex of the B-pocket is hydrophilic because of the Ser67 residue. The P9 peptide residue binds to the F-pocket in HLA-B*1501. The residues most important for the specificity of this pocket are Tyr74, Leu81, Leu95, Tyr123 and Trp147. These residues create a hydrophobic interior in the F-pocket and their spatial arrangement makes the pocket capable of containing large, bulky peptide side chains. Ser116 is located at the bottom of the F-pocket and makes the bottom of this pocket hydrophilic. Ser116, may act as a hydrogen-bonding partner and as such is a perfect place for binding of a Tyr9 peptide residue. Thus, based on structure information it is now possible to explain the peptide sequence specificity of HLA-B*1501 as previously determined by peptide binding and pool sequencing experiments.
- Institute of Medical Microbiology and Immunology, Panum Institute 18.3.20, Blegdamsvej 3B, 2200N Copenhagen, Denmark. g.roder@immi.ku.dk
Organizational Affiliation: