Selective Estrogen Receptor Modulators with Conformationally Restricted Side Chains. Synthesis and Structure-Activity Relationship of ERalpha-Selective Tetrahydroisoquinoline Ligands
Renaud, J., Bischoff, S.F., Buhl, T., Floersheim, P., Fournier, B., Geiser, M., Halleux, C., Kallen, J., Keller, H.J., Ramage, P.(2005) J Med Chem 48: 364-379
- PubMed: 15658851
- DOI: https://doi.org/10.1021/jm040858p
- Primary Citation of Related Structures:
1XQC - PubMed Abstract:
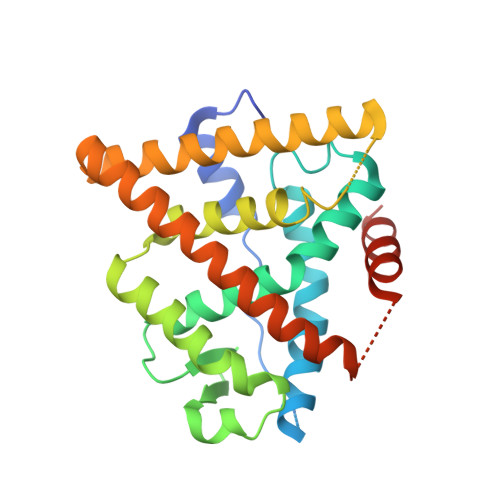
We disclose herein the discovery of estrogen receptor alpha (ERalpha) selective estrogen receptor modulators (SERMs) of the tetrahydroisoquinoline series that incorporate novel conformationally restricted side chains as replacement of the aminoethoxy residue typical of SERMs. Molecular modeling studies used in conjunction with the X-ray crystal structure of the ERalpha ligand binding domain (LBD) with raloxifene (7) suggested a diazadecaline moiety as a viable mimic of the SERM side chain. On the basis of this knowledge, the piperidinylethoxy moiety of our lead compound 60 was replaced by a diazadecaline subunit, providing the novel tetrahydroisoquinoline 29. In addition to exhibiting a binding affinity to ERalpha and antagonistic properties in the estrogen response element and MCF-7 assays similar to those of the parent compound 60, ligand 29 showed a reduced agonist behavior in the MCF-7 assay in the absence of 17beta-estradiol. These data point toward the fact that 29 may have a potential for breast cancer prevention/treatment in vivo, a feature which is particularly attractive in the quest for safe alternatives to hormone replacement therapy. In a pharmacokinetic experiment carried out in rats, 29 displayed an interesting profile, with a bioavailability of 49%. We also disclose the X-ray crystal structure of 29 in complex with ERalpha-LBD, which reveals the preferred configurations of 29 at the two chiral centers and the details of its interactions with the receptor. Finally, our structure-activity relationship studies show that other analogues bearing constrained side chains retain potency and antagonist activity and that a 3-OH substituted phenyl D-ring increases the selectivity of a set of piperazinyl-containing ligands in favor of ERalpha over ERbeta.
- Novartis Institute for Biomedical Research, WKL-136.6.81, CH-4002 Basel, Switzerland. johanne.renaud@pharma.novartis.com
Organizational Affiliation: