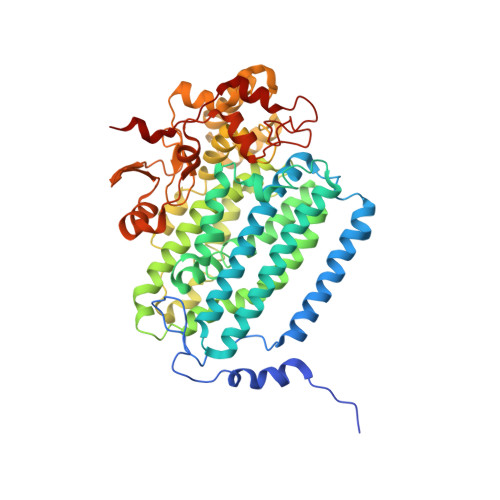
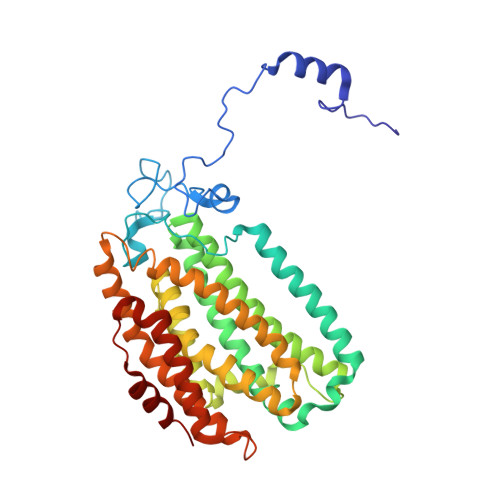
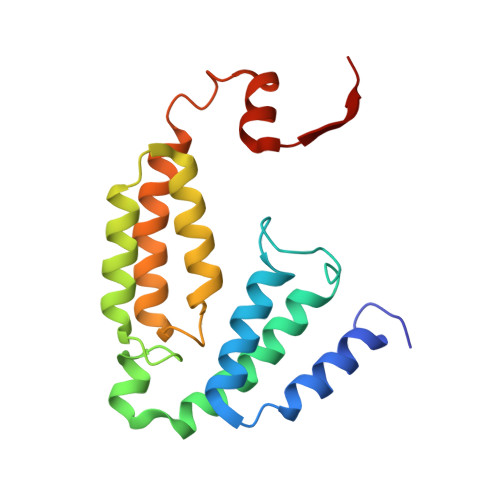
Preparation and X-ray Structures of Metal-Free, Dicobalt and Dimanganese Forms of Soluble Methane Monooxygenase Hydroxylase from Methylococcus capsulatus (Bath)
Sazinsky, M.H., Merkx, M., Cadieux, E., Tang, S., Lippard, S.J.(2004) Biochemistry 43: 16263-16276
- PubMed: 15610020 Search on PubMed
- DOI: https://doi.org/10.1021/bi048140z
- Primary Citation Related Structures:
1XMF, 1XMG, 1XMH - PubMed Abstract:
A three-component soluble methane monooxygenase (sMMO) enzyme system catalyzes the hydroxylation of methane to methanol at a carboxylate-bridged diiron center housed in the alpha-subunit of the hydroxylase (MMOH). Catalysis is facilitated by the presence of a regulatory protein (MMOB) and inhibited by MMOD, a protein of unknown function encoded in the sMMO operon. Both MMOB and MMOD are presumed to bind to the same region of the MMOH alpha-subunit. A colorimetric method for monitoring removal of Fe(II) from MMOH was developed using 1,10-phenanthroline and yields apo MMOH with <0.1 Fe/homodimer. With the use of this method, it was possible to investigate the X-ray structure of the apoenzyme and to perform metal reconstitution studies. Using MMOH from Methylococccus capsulatus (Bath), the effects of MMOB and MMOD on metal binding were studied and structural perturbations relevant to the function of this enzyme were identified. X-ray crystal structures of the apo, Mn(II)-soaked, and Co(II)-grown MMOH, determined to 2.3 A or greater resolution, reveal that the presence of metal ions is essential for the proper folding of helices E, F, and H of the alpha-subunit. The active sites of Mn(II)-soaked and Co(II)-grown MMOH are similar to that of reduced, native MMOH with notable differences in the metal-metal distances and ligand coordination sphere that may reflect how this dinuclear metal center might change in the presence of MMOB. MMOB and MMOD decrease the rate of removal of Fe(II) from the enzyme by 22- and 16-fold, respectively. On the basis of previous studies, it is hypothesized that MMOB, and perhaps MMOD, function to block solvent access to the MMOH active site. Finally, ITC studies and the observed disorder in helices E, F, and H in the apo and Mn(II)-soaked structures suggest that these regions of MMOH are critical for MMOB and MMOD binding.
- Department of Chemistry, Massachusetts Institute of Technology, Cambridge, Massachusetts 02139, USA.
Organizational Affiliation: