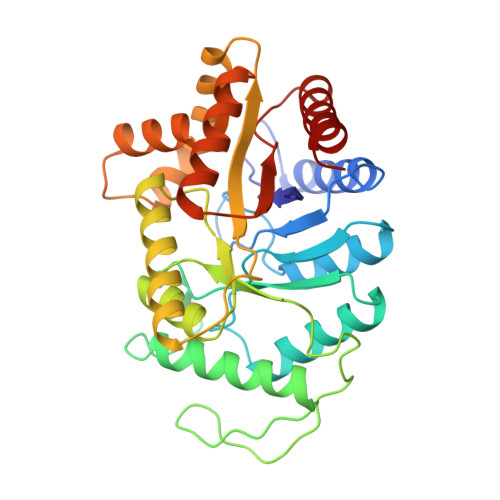
Structure of bacterial luciferase beta 2 homodimer: implications for flavin binding.
Tanner, J.J., Miller, M.D., Wilson, K.S., Tu, S.C., Krause, K.L.(1997) Biochemistry 36: 665-672
- PubMed: 9020763 Search on PubMed
- DOI: https://doi.org/10.1021/bi962511x
- Primary Citation Related Structures:
1XKJ - PubMed Abstract:
The crystal structure of the beta 2 homodimer of Vibrio harveyi luciferase has been determined to 2.5 A resolution by molecular replacement. Crystals were grown serendipitously using the alpha beta form of the enzyme. The subunits of the homodimer share considerable structural homology to the beta subunit of the alpha beta luciferase heterodimer. The four C-terminal residues that are disordered in the alpha beta structure are fully resolved in our structure. Four peptide bonds have been flipped relative to their orientations in the beta subunit of the alpha beta structure. The dimer interface of the homodimer is smaller than the interface of the heterodimer in terms of buried surface area and number of hydrogen bonds and salt links. Inspection of the subunits of our structure suggests that FMNH2 cannot bind to the beta 2 enzyme at the site that has been proposed for the alpha beta enzyme. However, we do uncover a potential FMNH2 binding pocket in the dimer interface, and we model FMN into this site. This proposed flavin binding motif is consistent with several lines of biochemical and structural evidence and leads to several conclusions. First, only one FMNH2 binds per homodimer. Second, we predict that reduced FAD and riboflavin should be poor substrates for beta 2. Third, the reduced activity of beta 2 compared to alpha beta is due to solvent exposure of the isoalloxazine ring in the beta 2 active site. Finally, we raise the question of whether our proposed flavin binding site could also be the binding site for flavin in the alpha beta enzyme.
- Department of Biochemical and Biophysical Sciences, University of Houston, Texas 77204-5934, USA.
Organizational Affiliation: