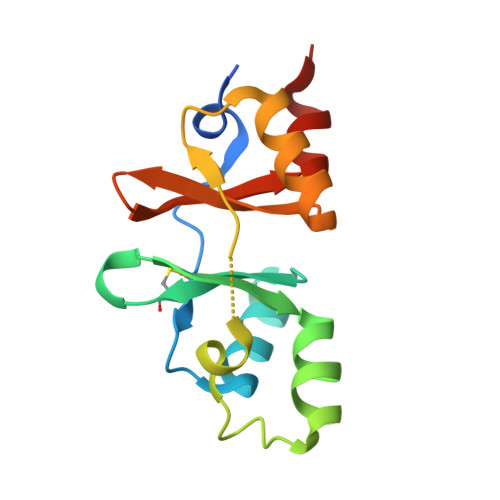
The structure and unusual protein chemistry of hypoxic response protein 1, a latency antigen and highly expressed member of the DosR regulon in Mycobacterium tuberculosis
Sharpe, M.L., Gao, C., Kendall, S.L., Baker, E.N., Lott, J.S.(2008) J Mol Biology 383: 822-836
- PubMed: 18640126 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.07.001
- Primary Citation Related Structures:
1XKF, 1Y5H - PubMed Abstract:
Mycobacterium tuberculosis adapts to cellular stresses such as decreased oxygen concentration, at least in part, by upregulation of the dormancy survival regulon, which is thought to be important for the bacterium's ability to enter a persistent state in its human host. We have determined the structure of hypoxic response protein 1, a protein encoded by one of the most strongly upregulated genes in the dormancy survival regulon. Hypoxic response protein 1 is an example of a 'cystathionine-beta-synthase-domain-only' protein; however, unlike other cystathionine-beta-synthase domains, it does not appear to bind AMP. The protein is proteolytically sensitive at its C-terminus and contains two unexpected disulfide bonds, one of which appears resistant to reducing agents in solution and is, therefore, most likely buried in the protein and is not solvent-accessible. We show that the protein is secreted from the bacterium in hypoxic in vitro culture and does not accumulate in the bacterial cell wall. The biological function of the protein remains unclear, but we suggest that it may contribute to the modulation of the host immune response. The work reported advances our understanding of the chemistry and cell biology of this intriguing and potentially important protein, and establishes a structural framework for future functional and immunological studies.
- School of Biological Sciences and Maurice Wilkins Center for Molecular Biodiscovery, University of Auckland, Private Bag 92109, Auckland 1142, New Zealand.
Organizational Affiliation: