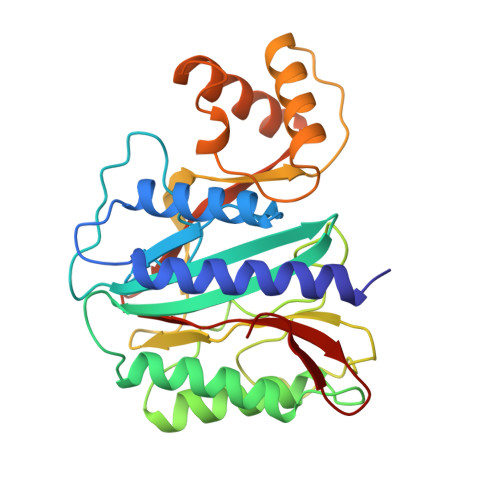
Crystal structure of methionine aminopeptidase from hyperthermophile, Pyrococcus furiosus.
Tahirov, T.H., Oki, H., Tsukihara, T., Ogasahara, K., Yutani, K., Ogata, K., Izu, Y., Tsunasawa, S., Kato, I.(1998) J Mol Biology 284: 101-124
- PubMed: 9811545 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.2146
- Primary Citation Related Structures:
1XGM, 1XGN, 1XGO, 1XGS - PubMed Abstract:
The structure of methionine aminopeptidase from hyperthermophile Pyrococcus furiosus (PfMAP) with an optimal growth temperature of 100 degreesC was determined by the multiple isomorphous replacement method and refined in three different crystal forms, one monoclinic and two hexagonal, at resolutions of 2.8, 2.9, and 3.5 A. The resolution of the monoclinic crystal form was extended to 1.75 A by water-mediated transformation to a low-humidity form, and the obtained diffraction data used for high-resolution structure refinement. This is the first description of a eukaryotic type methionine aminopeptidase structure. The PfMAP molecule is composed of two domains, a catalytic domain and an insertion domain, connected via two antiparallel beta-strands. The catalytic domain, which possesses an internal 2-fold symmetry and contains two cobalt ions in the active site, resembles the structure of a prokaryotic type MAP from Escherichia coli (EcMAP), while the structure of the insertion domain containing three helices has a novel fold and accounts for a major difference between the eukaryotic and prokaryotic types of methionine aminopeptidase. Analysis of the PfMAP structure in comparison with EcMAP and other mesophile proteins reveals several factors which may contribute to the hyperthermostability of PfMAP: (1) a significantly high number of hydrogen bonds and ion-pairs between side-chains of oppositely charged residues involved in the stabilization of helices; (2) an increased number of hydrogen bonds between the positively charged side-chain and neutral oxygen; (3) a larger number of buried water molecules involved in crosslinking the backbone atoms of sequentially separate segments; (4) stabilization of two antiparallel beta-strands connecting the two domains of the molecule by proline residues; (5) shortening of N and C-terminal tails and stabilization of the loop c3E by deletion of three residues.
- Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita, Osaka, 565, Japan.
Organizational Affiliation: