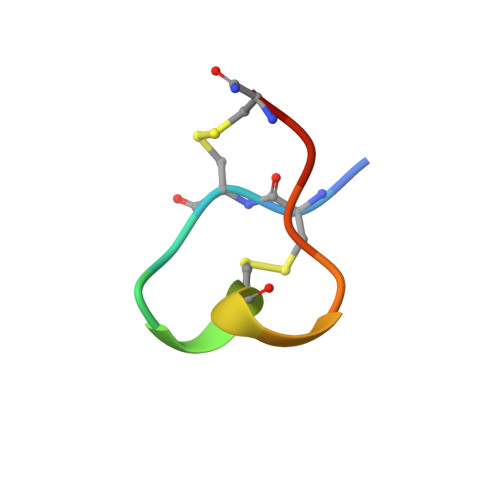
Structure determination of the three disulfide bond isomers of alpha-conotoxin GI: a model for the role of disulfide bonds in structural stability.
Gehrmann, J., Alewood, P.F., Craik, D.J.(1998) J Mol Biology 278: 401-415
- PubMed: 9571060 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1998.1701
- Primary Citation Related Structures:
1XGA, 1XGB, 1XGC - PubMed Abstract:
The three possible disulfide bonded isomers of alpha-conotoxin GI have been selectively synthesised and their structures determined by 1H NMR spectroscopy. alpha-Conotoxin GI derives from the venom of Conus geographus and is a useful neuropharmacological tool as it selectively binds to the nicotinic acetylcholine receptor (nAChR), a ligand-gated ion channel involved in nerve signal transmission. The peptide has the sequence ECCNPACGRHYSC-NH2, and the three disulfide bonded isomers are referred to as GI(2-7;3-13), GI(2-13;3-7) and GI(2-3;7-13). The NMR structure for the native isomer GI(2-7;3-13) is of excellent quality, with a backbone pairwise RMSD of 0.16 A for a family of 35 structures, and comprises primarily a distorted 310 helix between residues 5 to 11. The two non-native isomers exhibit multiple conformers in solution, with the major populated forms being different in structure both from each other and from the native form. Structure-activity relationships for the native GI(2-7;3-13) as well as the role of the disulfide bonds on folding and stability of the three isomers are examined. It is concluded that the disulfide bonds in alpha-conotoxin GI play a crucial part in determining both the structure and stability of the peptide. A trend for increased conformational heterogeneity was observed in the order of GI(2-7;3-13)
- Centre for Drug Design and Development, University of Queensland, Brisbane, QLD 4072, Australia.
Organizational Affiliation: