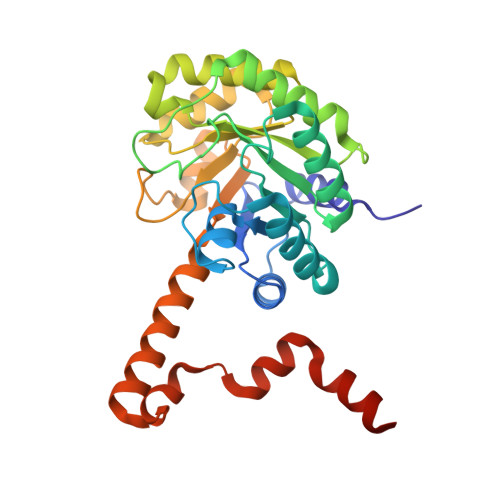
Crystal Structures of 2-Methylisocitrate Lyase in Complex with Product and with Isocitrate Inhibitor Provide Insight into Lyase Substrate Specificity, Catalysis and Evolution
Liu, S., Lu, Z., Han, Y., Melamud, E., Dunaway-Mariano, D., Herzberg, O.(2005) Biochemistry 44: 2949-2962
- PubMed: 15723538
- DOI: https://doi.org/10.1021/bi0479712
- Primary Citation Related Structures:
1OQF, 1XG3, 1XG4 - PubMed Abstract:
Two crystal structures of the C123S mutant of 2-methylisocitrate lyase have been determined, one with the bound reaction products, Mg(2+)-pyruvate and succinate, and the second with a bound Mg(2+)-(2R,3S)-isocitrate inhibitor. Comparison with the structure of the wild-type enzyme in the unbound state reveals that the enzyme undergoes a conformational transition that sequesters the ligand from solvent, as previously observed for two other enzyme superfamily members, isocitrate lyase and phosphoenolpyruvate mutase. The binding modes reveal the determinants of substrate specificity and stereoselectivity, and the stringent specificity is verified in solution using various potential substrates. A model of bound 2-methylisocitrate has been developed based on the experimentally determined structures. We propose a catalytic mechanism involving an alpha-carboxy-carbanion intermediate/transition state, which is consistent with previous stereochemical experiments showing inversion of configuration at the C(3) of 2-methylisocitrate. Structure-based sequence analysis and phylogenic tree construction reveal determinants of substrate specificity, highlight nodes of divergence of families, and predict enzyme families with new functions.
- Center for Advanced Research in Biotechnology, University of Maryland Biotechnology Institute, Rockville, Maryland 20850, USA.
Organizational Affiliation: